About This Guide
This guide explains SEC concepts from a practical perspective—what
you need to know to use measure.sec effectively. It’s not a
textbook; every concept ties back to package functionality with guidance
on when and why to use different approaches.
Who this is for: Users with a chemistry background who are new to SEC or want to understand when to use which package features.
How to use it: Read straight through for a conceptual foundation, or jump to specific sections when you need decision guidance.
How SEC Data Flows Through measure.sec
The Processing Pipeline
SEC data processing follows a consistent pattern: raw detector signals are converted to a common format, processed to remove artifacts, calibrated to molecular weight, and then used to calculate results. Understanding this flow helps you build recipes that make sense.
#> Warning in geom_segment(aes(x = 1.15, xend = 4.85, y = 1, yend = 1), arrow = arrow(length = unit(0.3, : All aesthetics have length 1, but the data has 5 rows.
#> ℹ Please consider using `annotate()` or provide this layer with data containing
#> a single row.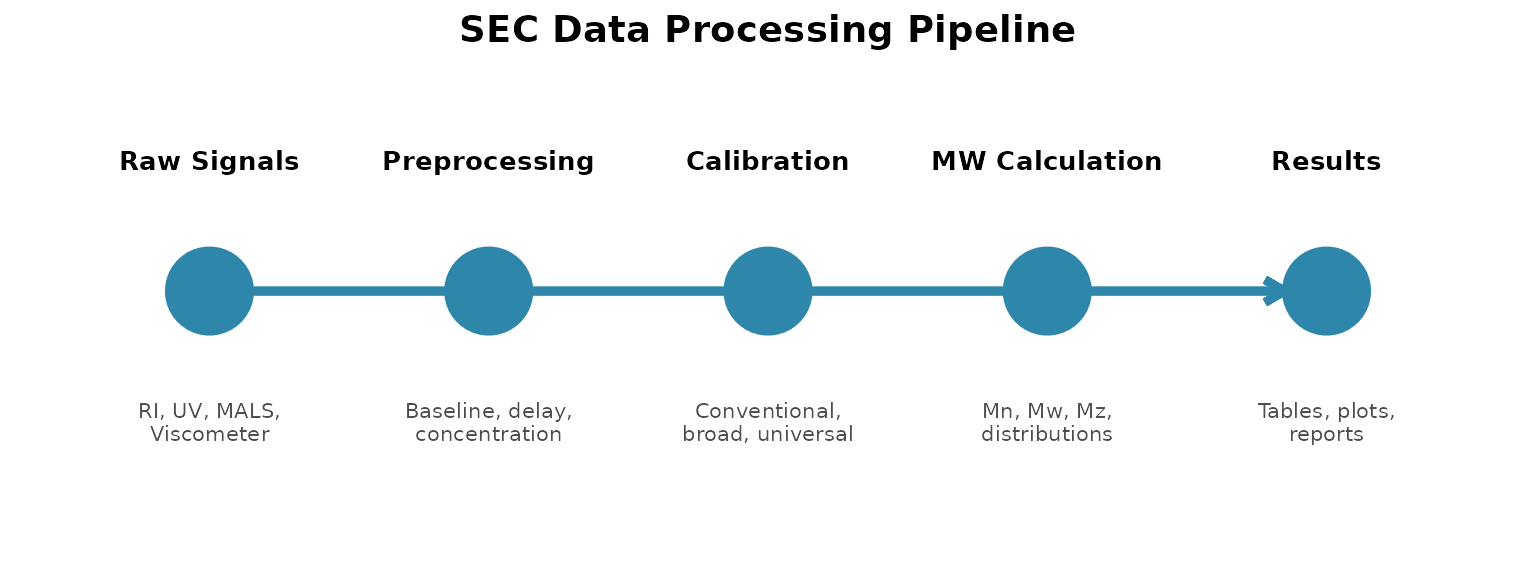
Stage 1: Input Signals
Every recipe starts by converting raw detector signals into
measure format using
step_measure_input_long(). This standardizes different
detectors into a common structure where each measurement has a location
(elution time or volume) and a value (signal intensity).
recipe(...) |>
step_measure_input_long(ri_signal, location = vars(elution_time), col_name = "ri") |>
step_measure_input_long(uv_signal, location = vars(elution_time), col_name = "uv")Stage 2: Preprocessing
Before calibration, signals need cleaning:
| Step | What It Does | When to Use |
|---|---|---|
step_sec_detector_delay() |
Aligns multi-detector signals | Always with multiple detectors |
step_sec_baseline() |
Removes baseline drift | Almost always |
step_sec_ri() / step_sec_uv()
|
Normalizes by response factor | Before concentration calculation |
step_sec_concentration() |
Converts to mass concentration | Before light scattering or viscometry |
step_sec_band_broadening() |
Corrects peak spreading | High-resolution MW distributions |
Order matters: Detector delay correction must come first. Baseline correction should precede normalization. Concentration is needed before MALS calculations.
Stage 3: Calibration
Calibration converts elution time to molecular weight. The right approach depends on your standards and goals:
| Method | Function | Best For |
|---|---|---|
| Conventional | step_sec_conventional_cal() |
Narrow standards, same polymer type |
| Broad standard | step_sec_broad_standard() |
Single characterized reference |
| Universal | step_sec_universal_cal() |
Different polymer types |
| Absolute (MALS) | step_sec_mals() |
True MW, no standards needed |
Stage 4: MW Calculation
With calibration complete, calculate molecular weight results:
| Step | Output |
|---|---|
step_sec_mw_averages() |
Mn, Mw, Mz, dispersity |
step_sec_mw_distribution() |
d(W)/d(log M) curve |
step_sec_mw_fractions() |
Cumulative fractions at MW cutoffs |
Stage 5: Export and Visualization
Extract results for reporting:
-
measure_sec_summary_table(): Publication-ready summary statistics -
measure_sec_slice_table(): Point-by-point data for custom analysis -
plot_sec_*()family: Chromatograms, MWDs, calibration curves
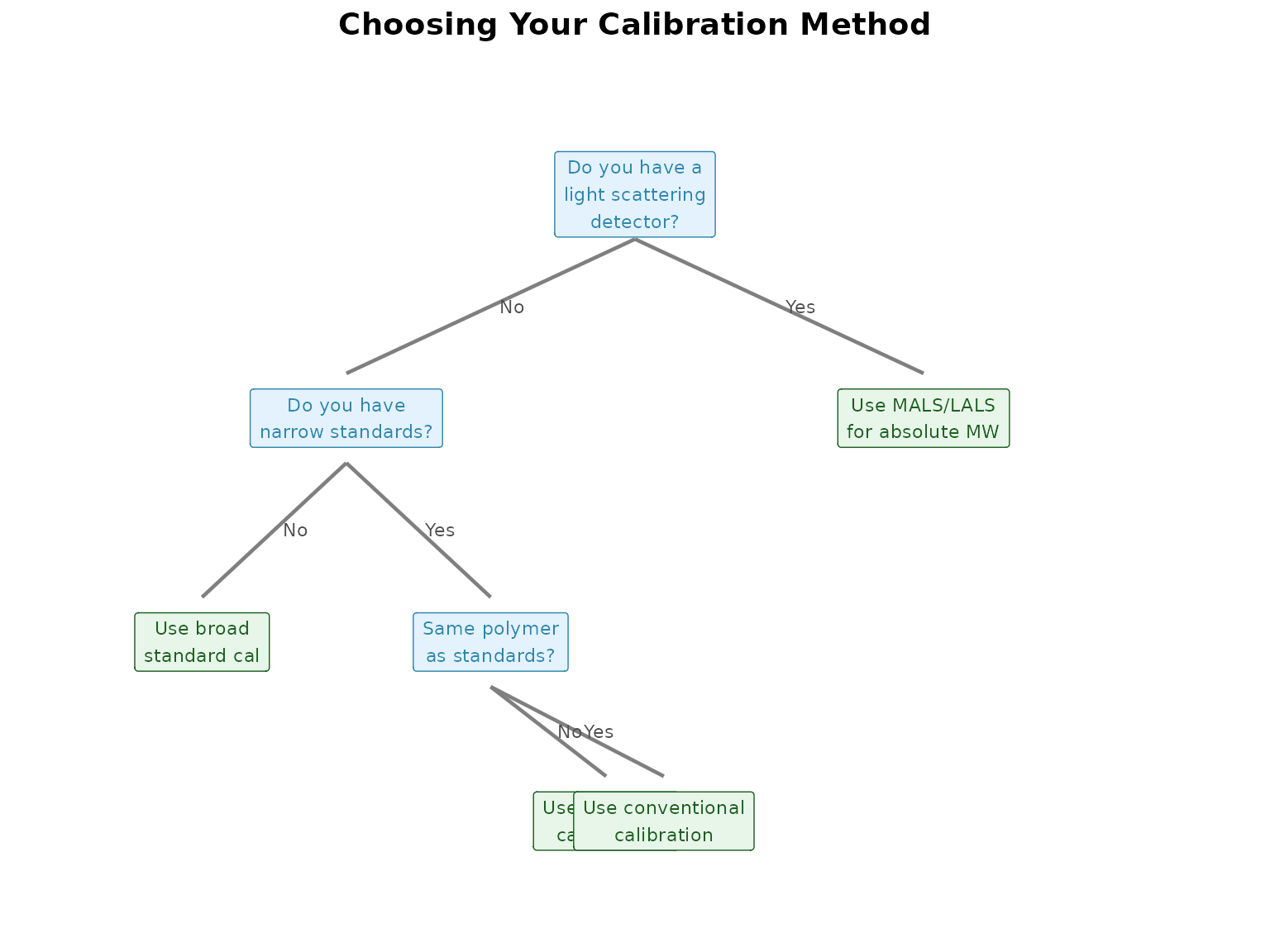
Choosing Your Calibration Approach
Calibration is where most users face their first major decision. This section helps you choose the right approach.
Conventional Calibration
When to use: You have narrow MW standards (low dispersity) of the same polymer type as your samples.
How it works: Fit a polynomial relationship between log(MW) and elution time using standards. Apply this curve to unknown samples.
step_sec_conventional_cal(
standards_data = standards,
degree = 3 # Polynomial order (usually 3)
)Advantages: Simple, well-understood, widely used. Calibration curve provides diagnostics (R2, residuals).
Limitations: Assumes your sample behaves like the standards. A branched polymer will report incorrect MW because it elutes later than a linear polymer of the same MW.
Best practices:
- Use 5+ standards spanning your MW range of interest
- Check residuals—systematic patterns indicate poor fit
- Don’t extrapolate far beyond your calibration range
Broad Standard Calibration
When to use: You have one well-characterized broad-distribution reference material instead of multiple narrow standards.
How it works: Uses the known MWD of the reference to establish the MW-elution relationship in a single injection.
step_sec_broad_standard(
reference_mwd = known_distribution,
reference_mn = 50000,
reference_mw = 150000
)Advantages: Single injection, good for routine QC. Convenient when narrow standards aren’t available.
Limitations: Accuracy depends entirely on how well you know the reference material’s MWD. Any error in the reference propagates to all samples.
Universal Calibration
When to use: Your samples are a different polymer type than your available standards.
How it works: Calibrates on hydrodynamic volume ([η] × M) rather than molecular weight directly. Uses Mark-Houwink parameters to convert between polymer types.
step_sec_universal_cal(
# Calibration standards (e.g., polystyrene)
reference_k = 1.14e-4,
reference_a = 0.716,
# Your sample (e.g., PMMA)
sample_k = 6.0e-5,
sample_a = 0.73
)Requires: Intrinsic viscosity data (from viscometer) AND Mark-Houwink parameters for both polymer types.
Advantages: Accurate MW for different polymer types using common standards.
Limitations: You need reliable K and a values for your sample, which may require literature search or independent measurement. Doesn’t work well for highly branched polymers where Mark-Houwink parameters aren’t constant.
Absolute MW (Light Scattering)
When to use: You need true molecular weight independent of polymer structure, or have no appropriate standards.
How it works: Light scattering intensity is directly proportional to molecular weight × concentration. No column calibration needed.
step_sec_mals(
dn_dc_column = "dn_dc" # Refractive index increment
)Requires: Light scattering detector (MALS, LALS, or RALS), accurate dn/dc value, good signal quality.
Advantages: True absolute MW. Works for any polymer type. Reveals branching through Rg-MW relationships. Required for proteins and complex polymers.
Limitations: More expensive instrumentation. Sensitive to dust and bubbles. Low MW samples may have weak scattering signal.
Understanding Your Detectors
Each detector type provides different information. Knowing what each measures helps you interpret results and troubleshoot problems.
Concentration Detectors
These tell you how much material is present at each elution slice.
Refractive Index (RI)
What it measures: Change in refractive index proportional to concentration.
Key parameter: dn/dc (refractive index increment, mL/g)
step_sec_ri(dn_dc_column = "dn_dc")| Pros | Cons |
|---|---|
| Universal—responds to all polymers | Sensitive to temperature |
| Mass-based response | Sensitive to mobile phase composition |
| Most common detector | Baseline drift common |
When to use: Default concentration detector for most polymer SEC. Essential for MW calculations.
UV Detector
What it measures: Absorbance at specific wavelength(s).
Key parameter: Extinction coefficient (ε, mL/(g·cm))
step_sec_uv(extinction_column = "extinction_coef", wavelength = 280)| Pros | Cons |
|---|---|
| Very sensitive | Only detects UV-active chromophores |
| Selective | Response depends on chromophore content |
| Stable baseline | May need multiple wavelengths |
When to use: Proteins (280 nm), aromatic polymers, or when combined with RI for composition analysis.
DAD (Diode Array)
What it measures: Full UV-Vis spectrum at each point.
step_sec_dad(wavelengths = c(254, 280, 320))When to use: Complex samples where spectral information helps identify components. Copolymer composition with wavelength-dependent absorption.
Molecular Weight Detectors
These measure MW directly, independent of calibration.
Multi-Angle Light Scattering (MALS)
What it measures: Scattered light intensity at multiple angles.
Provides: Absolute MW and radius of gyration (Rg) at each slice.
step_sec_mals(dn_dc_column = "dn_dc")| Best for | Limitations |
|---|---|
| Large molecules (> 10 kDa) | Requires very clean samples |
| Rg measurement | Signal weak for small molecules |
| Branching analysis | Expensive |
See also: MALS Detection vignette
Low-Angle Light Scattering (LALS)
What it measures: Scattered light near 0° angle.
step_sec_lals(dn_dc_column = "dn_dc")| Best for | Limitations |
|---|---|
| Medium molecules | No Rg measurement |
| Simpler than MALS | Less accurate for large molecules |
| Lower cost | Still needs clean samples |
Right-Angle Light Scattering (RALS)
What it measures: Scattered light at 90°.
step_sec_rals(dn_dc_column = "dn_dc")| Best for | Limitations |
|---|---|
| Small molecules | Biased for large molecules |
| Screening/QC | No structural information |
| Lowest cost LS option | Limited to narrow MW range |
See also: LALS/RALS Detection vignette
Hydrodynamic Detectors
These measure molecular size and conformation in solution.
Viscometer
What it measures: Differential pressure across a capillary, related to solution viscosity.
Provides: Intrinsic viscosity [η] when combined with concentration.
step_sec_viscometer()
step_sec_intrinsic_visc(visc_col = "visc", conc_col = "concentration")| Best for | Limitations |
|---|---|
| Universal calibration | Requires concentration data |
| Branching characterization | Sensitive to bubbles |
| Mark-Houwink parameters | Additional complexity |
Common Pitfalls and Solutions
This section addresses problems you’re likely to encounter and how
measure.sec helps you fix them.
Baseline Issues
Symptoms: Sloped baseline, wavy baseline, or baseline that doesn’t return to zero.
Causes: Temperature drift (RI), mobile phase mixing, column equilibration, late-eluting contaminants.
Solution: Use step_sec_baseline() with
an appropriate method:
step_sec_baseline(
measures = "ri",
method = "linear", # Simple start-to-end connection
# or
method = "spline", # Follows gradual curves
# or
method = "asymmetric" # Handles uneven noise
)Which method to choose:
- linear: Fast, works when baseline is truly straight
- spline: Better for gradual drift or temperature effects
- asymmetric: Best for noisy data or when peak tails affect baseline
Inter-Detector Delay
Symptoms: Calculated MW varies with injection volume, or MW appears to vary across a peak that should be uniform.
Cause: Detectors in series experience different delays. A 150 µL delay volume at 1 mL/min means 9 seconds of offset—which can shift MW by orders of magnitude.
Solution: Use step_sec_detector_delay()
before any MW calculations:
step_sec_detector_delay(
reference = "ri",
delay_volumes = c(uv = -0.05, mals = 0.15) # mL
)How to measure delays:
- Inject a low-MW marker (toluene, polymer standard)
- Record peak apex time on each detector
- Convert time difference to volume: delay = Δt × flow_rate
Tip: Re-measure after column changes or major maintenance.
Band Broadening
Symptoms: MWD appears broader than expected. High MW tail or low MW shoulder.
Cause: Peak spreading from extra-column volume, column dispersion, or injection effects.
Solution: Use
step_sec_band_broadening() to deconvolve the true
distribution:
step_sec_band_broadening(
sigma = 0.03 # Peak spreading parameter (mL)
)Estimate sigma using a narrow standard—the peak width should be entirely from broadening:
sigma <- estimate_sigma(
narrow_standard_data,
signal_col = "ri"
)Out-of-Range Calibration
Symptoms: Extrapolated MW values seem unreasonable. Very high or very low MW fractions.
Cause: Sample contains material outside the MW range of your calibration standards.
Solution: Options in
step_sec_conventional_cal():
step_sec_conventional_cal(
standards_data = standards,
extrapolate = "linear", # Continue slope beyond range
# or
extrapolate = "constant", # Clamp at boundary
# or
extrapolate = "none" # Return NA outside range
)Best practice: Use standards that span your sample’s MW range. If extrapolation is unavoidable, use “linear” with caution and flag those results.
Noisy Light Scattering
Symptoms: Jagged LS signal, especially at low concentration regions.
Causes: Dust, bubbles, low sample concentration, detector noise.
Solutions:
- Filter samples: 0.22 µm minimum, 0.1 µm for MALS
- Degas mobile phase: Remove dissolved gases
- Increase concentration: More sample = more scattering signal
- Apply smoothing: In step parameters or post-processing
Interpreting Results
Molecular Weight Averages
SEC produces multiple MW averages. Each tells you something different:
| Average | Formula | What It Emphasizes |
|---|---|---|
| Mn | Σ(niMi) / Σni | Number of chains—sensitive to low MW |
| Mw | Σ(niMi2) / Σ(niMi) | Mass of chains—sensitive to high MW |
| Mz | Σ(niMi3) / Σ(niMi2) | Very sensitive to high MW tail |
Practical interpretation:
- Mn tells you about the most numerous species
- Mw tells you where most of the mass is
- Mz warns about high MW contamination or aggregation
Dispersity (Đ)
Dispersity = Mw/Mn measures the breadth of the distribution.
| Đ Value | Interpretation |
|---|---|
| 1.0 | Perfectly monodisperse (theoretical limit) |
| < 1.1 | Very narrow (controlled polymerization) |
| 1.5 - 2.0 | Typical for step-growth or conventional free radical |
| > 3.0 | Very broad, may indicate blending or degradation |
Tip: If Đ < 1.0, something is wrong—often a calibration or integration problem.
When Results Look Wrong
MW seems too high or too low:
- Check calibration: Are standards correctly identified?
- Check detector delays: Misalignment shifts apparent MW
- Check dn/dc: Wrong value directly scales MW from LS
Dispersity seems too narrow:
- Check integration limits: Too narrow excludes real distribution
- Check band broadening: Unaccounted broadening inflates apparent width
- Sample may genuinely be narrow
MW varies between injections:
- Check baseline reproducibility
- Check column equilibration
- Check sample stability (aggregation, degradation)
Negative MW at distribution edges:
- Normal at very low signal—noise can go negative
- Set appropriate thresholds in MW calculation steps
- Focus on the region with good signal-to-noise
See Also
- Getting Started - Your first SEC analysis
- Comprehensive SEC Analysis - Full reference for all features
- Multi-Detector SEC - Detailed multi-detector workflows
- Calibration Management - Saving and reusing calibrations