Overview
This guide shows you how to save, load, and reuse SEC calibrations with measure.sec. Key tasks covered:
- Save a calibration from a prepped recipe
- Load and reuse a saved calibration
- Inspect calibration quality
- Compare calibrations over time
Setup
library(measure)
#> Loading required package: recipes
#> Loading required package: dplyr
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
#>
#> Attaching package: 'recipes'
#> The following object is masked from 'package:stats':
#>
#> step
library(measure.sec)
library(recipes)
library(dplyr)
library(ggplot2)How to Create and Save a Calibration
Step 1: Fit a Calibration
First, create a recipe with calibration standards and prep it to fit the calibration:
# Load polystyrene standards
data(sec_ps_standards)
# View the standards
sec_ps_standards |>
select(standard_name, mp, log_mp, retention_time) |>
head(8)
#> # A tibble: 8 × 4
#> standard_name mp log_mp retention_time
#> <chr> <dbl> <dbl> <dbl>
#> 1 PS-3150000 3150000 6.50 11.2
#> 2 PS-1870000 1870000 6.27 11.6
#> 3 PS-1090000 1090000 6.04 12.1
#> 4 PS-630000 630000 5.80 12.6
#> 5 PS-430000 430000 5.63 13.2
#> 6 PS-216000 216000 5.33 13.8
#> 7 PS-120000 120000 5.08 14.3
#> 8 PS-67500 67500 4.83 15.0
# Prepare standards with required column names
# The step expects 'retention' and 'log_mw' columns
ps_cal_standards <- sec_ps_standards |>
select(retention = retention_time, log_mw = log_mp)
# Create a simple data frame for the recipe
# (In practice, this would be your chromatogram data)
sample_data <- tibble(
sample_id = "test",
elution_time = seq(10, 20, by = 0.01),
ri_signal = dnorm(seq(10, 20, by = 0.01), mean = 14, sd = 1)
)
# Create and prep a recipe with conventional calibration
rec <- recipe(~ ., data = sample_data) |>
step_measure_input_long(
ri_signal,
location = vars(elution_time),
col_name = "ri"
) |>
step_sec_conventional_cal(
standards = ps_cal_standards,
fit_type = "cubic"
)
# Prep the recipe (this fits the calibration)
prepped <- prep(rec)
#> Warning: Standard at 12.58 has 14.4% MW deviation.
#> ℹ Consider removing outlier standards or using a different fit type.
#> Warning: 115 points (11.5%) are outside calibration range.
#> ℹ Calibration range: 11.15 to 20.79Step 2: Save the Calibration
Use save_sec_calibration() to extract and save the
calibration for reuse:
# Create a temporary file for this example
cal_file <- tempfile(fileext = ".rds")
# Save with metadata for traceability
save_sec_calibration(
prepped,
cal_file,
metadata = list(
column = "PLgel 5um Mixed-C",
instrument = "Agilent 1260",
analyst = "JW",
notes = "Monthly calibration - January 2024"
)
)
#> ✔ Saved cubic calibration to /tmp/RtmpnCnUxt/file284c38a1b389.rdsFormat Options
Two formats are supported:
| Format | Extension | Best For |
|---|---|---|
| RDS | .rds |
Default. Preserves all R objects exactly |
| YAML | .yaml |
Human-readable, good for version control |
# Save as YAML for human readability
save_sec_calibration(prepped, "calibration.yaml")How to Load and Reuse a Calibration
Load a Saved Calibration
# Load the calibration
cal <- load_sec_calibration(cal_file)
#> ℹ Loaded cubic calibration (R\u00b2 = 0.9995)
# View calibration details
print(cal)
#>
#> ── SEC Calibration ─────────────────────────────────────────────────────────────
#> Fit type: cubic (degree 3)
#> Calibration range: 11.15 to 20.79
#> Standards: 16
#>
#> ── Fit Quality ──
#>
#> R\u00b2: 0.999466
#> RMSE (log MW): 0.0294
#> Max % deviation: 14.43%
#>
#> ── Settings ──
#>
#> Extrapolation: warn
#> Output column: mw
#> Log output: TRUE
#>
#> ── Metadata ──
#>
#> Created: 2026-02-02 17:16:55
#> measure.sec version: 0.0.0.9000
#>
#> ── User Metadata
#> column: PLgel 5um Mixed-C
#> instrument: Agilent 1260
#> analyst: JW
#> notes: Monthly calibration - January 2024Use in a New Recipe
Pass the loaded calibration to
step_sec_conventional_cal() to skip fitting:
# New sample data
new_samples <- tibble(
sample_id = "unknown_polymer",
elution_time = seq(10, 20, by = 0.01),
ri_signal = dnorm(seq(10, 20, by = 0.01), mean = 15, sd = 0.8)
)
# Create recipe using saved calibration (no fitting needed)
rec_new <- recipe(~ ., data = new_samples) |>
step_measure_input_long(
ri_signal,
location = vars(elution_time),
col_name = "ri"
) |>
step_sec_conventional_cal(calibration = cal)
# Prep and bake
prepped_new <- prep(rec_new)
#> ℹ Using pre-loaded cubic calibration (R\u00b2 = 0.9995)
#> Warning: 115 points (11.5%) are outside calibration range.
#> ℹ Calibration range: 11.15 to 20.79
result <- bake(prepped_new, new_data = NULL)How to Inspect Calibration Quality
View Fit Diagnostics
# Summary shows per-standard results
summary(cal)
#>
#> ── SEC Calibration ─────────────────────────────────────────────────────────────
#> Fit type: cubic (degree 3)
#> Calibration range: 11.15 to 20.79
#> Standards: 16
#>
#> ── Fit Quality ──
#>
#> R\u00b2: 0.999466
#> RMSE (log MW): 0.0294
#> Max % deviation: 14.43%
#>
#> ── Settings ──
#>
#> Extrapolation: warn
#> Output column: mw
#> Log output: TRUE
#>
#> ── Metadata ──
#>
#> Created: 2026-02-02 17:16:55
#> measure.sec version: 0.0.0.9000
#>
#> ── User Metadata
#> column: PLgel 5um Mixed-C
#> instrument: Agilent 1260
#> analyst: JW
#> notes: Monthly calibration - January 2024
#>
#> ── Per-Standard Results ──
#>
#> # A tibble: 16 × 10
#> location actual_log_mw predicted_log_mw residual_log_mw actual_mw
#> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 11.2 6.50 6.47 0.0294 3150000.
#> 2 11.6 6.27 6.28 -0.00364 1870000.
#> 3 12.1 6.04 6.06 -0.0192 1090000
#> 4 12.6 5.80 5.86 -0.0585 630000.
#> 5 13.2 5.63 5.59 0.0421 430000.
#> 6 13.8 5.33 5.32 0.0126 216000
#> 7 14.3 5.08 5.11 -0.0300 120000.
#> 8 15.0 4.83 4.79 0.0413 67500.
#> 9 15.5 4.53 4.55 -0.0262 33500.
#> 10 16.1 4.30 4.26 0.0352 19800
#> 11 16.7 3.99 4.01 -0.0197 9680
#> 12 17.4 3.70 3.70 0.00359 5030
#> 13 17.9 3.47 3.46 0.0146 2970
#> 14 18.9 3.02 3.02 0.00222 1050
#> 15 19.4 2.76 2.81 -0.0419 580
#> 16 20.8 2.21 2.19 0.0181 162
#> # ℹ 5 more variables: predicted_mw <dbl>, pct_deviation <dbl>,
#> # prediction_se <dbl>, ci_lower_log_mw <dbl>, ci_upper_log_mw <dbl>Key Quality Metrics
| Metric | Good Value | Meaning |
|---|---|---|
| R² | > 0.999 | Fit explains variance |
| RMSE (log MW) | < 0.05 | Prediction error in log units |
| Max % deviation | < 5% | Worst individual standard |
Visualize the Calibration Curve
# Plot calibration curve from standards data
plot_sec_calibration(sec_ps_standards)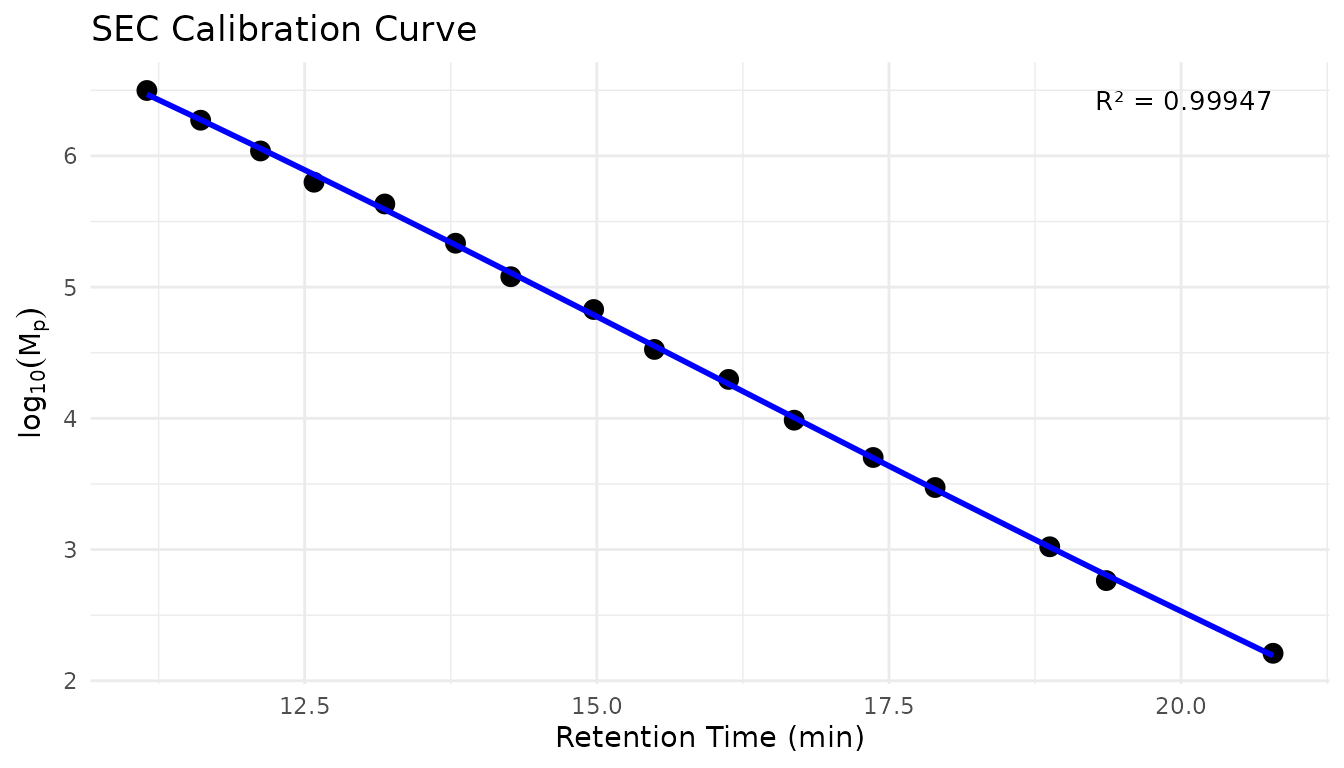
# With residuals panel to check for systematic errors
plot_sec_calibration(sec_ps_standards, show_residuals = TRUE)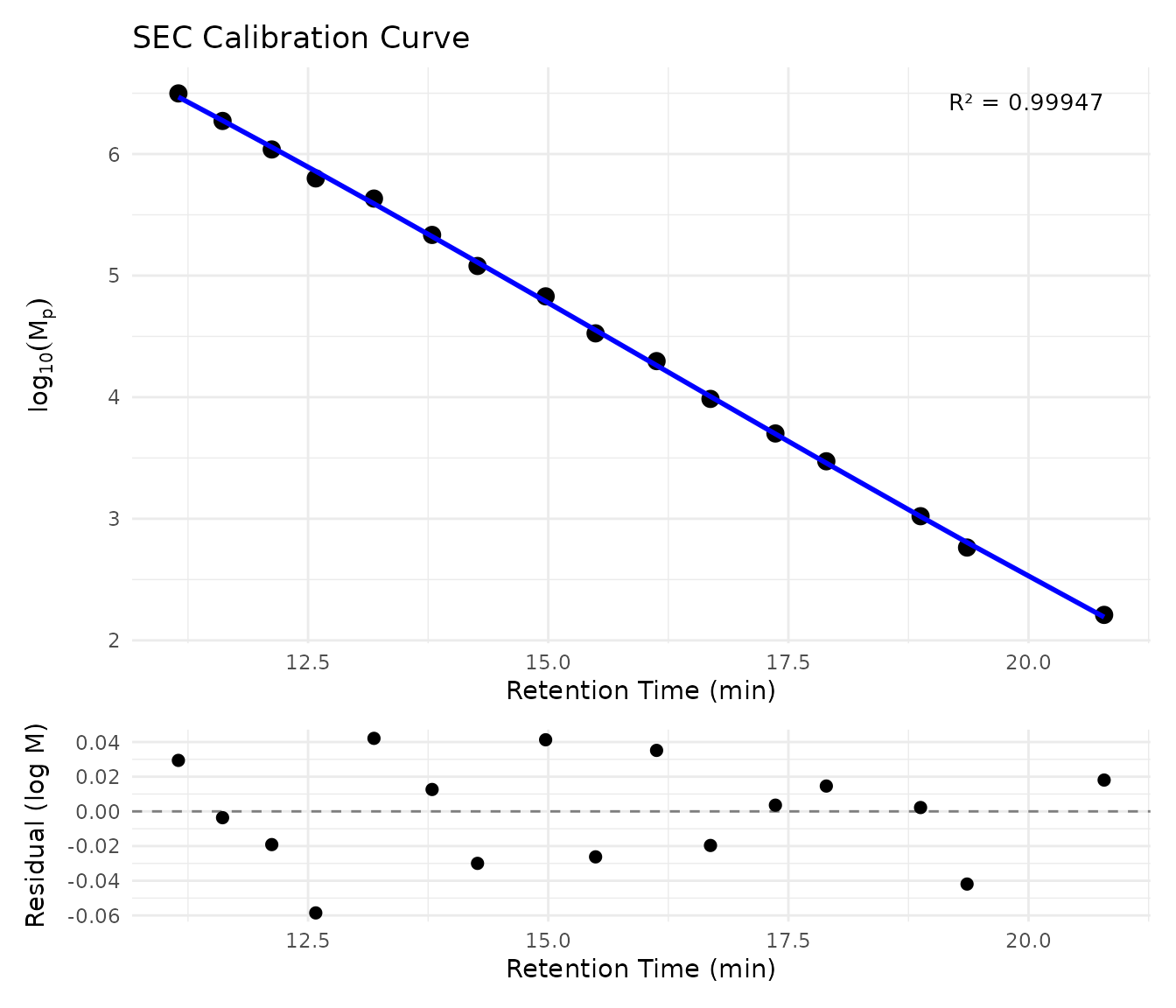
When to Recalibrate vs. Reuse
Best Practices
Naming Conventions
Use descriptive, consistent file names:
# Good: includes key information
"ps_plgel-mixed-c_2024-01-15.rds"
"pmma_superose6_instrument2_2024-01.yaml"
# Avoid: ambiguous names
"calibration.rds"
"cal_new.rds"Include Metadata
Always add metadata for traceability:
save_sec_calibration(
prepped,
"ps_calibration.rds",
metadata = list(
column = "PLgel 5um Mixed-C, S/N ABC123",
instrument = "Agilent 1260 #2",
analyst = "Jane Doe",
date = Sys.Date(),
mobile_phase = "THF + 0.1% BHT",
flow_rate = "1.0 mL/min",
temperature = "35C",
notes = "Monthly recalibration after column flush"
)
)Version Control (YAML Format)
For labs using git or similar version control:
# YAML format is diff-friendly
save_sec_calibration(prepped, "calibrations/ps_current.yaml")
# Git will show meaningful diffs when calibration changesLab Workflow Integration
A typical workflow:
- Monthly: Run calibration standards, save new calibration with date
- Daily: Load current calibration for sample analysis
- Archive: Keep old calibrations for historical reference
- Compare: Periodically compare calibrations to detect drift
# Monthly calibration routine
monthly_cal <- prep(calibration_recipe)
save_sec_calibration(
monthly_cal,
sprintf("calibrations/ps_%s.rds", format(Sys.Date(), "%Y-%m")),
metadata = list(analyst = Sys.info()["user"])
)
# Daily sample analysis
cal <- load_sec_calibration("calibrations/ps_2024-01.rds")
samples_recipe <- recipe(...) |>
step_sec_conventional_cal(calibration = cal) |>
prep()Cleanup
# Remove temporary file
unlink(cal_file)See Also
- Getting Started - Basic SEC workflow and concepts
- System Suitability - Validate column performance
- Multi-Detector SEC - Absolute MW without calibration
- Exporting Results - Summary tables and reports
Session Info
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.1 measure.sec_0.0.0.9000 measure_0.0.1.9002
#> [4] recipes_1.3.1 dplyr_1.1.4
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.56 bslib_0.10.0
#> [4] lattice_0.22-7 vctrs_0.7.1 tools_4.5.2
#> [7] generics_0.1.4 parallel_4.5.2 tibble_3.3.1
#> [10] pkgconfig_2.0.3 Matrix_1.7-4 data.table_1.18.2.1
#> [13] RColorBrewer_1.1-3 S7_0.2.1 desc_1.4.3
#> [16] lifecycle_1.0.5 compiler_4.5.2 farver_2.1.2
#> [19] textshaping_1.0.4 codetools_0.2-20 htmltools_0.5.9
#> [22] class_7.3-23 sass_0.4.10 yaml_2.3.12
#> [25] prodlim_2025.04.28 tidyr_1.3.2 pillar_1.11.1
#> [28] pkgdown_2.2.0 jquerylib_0.1.4 MASS_7.3-65
#> [31] cachem_1.1.0 gower_1.0.2 rpart_4.1.24
#> [34] parallelly_1.46.1 lava_1.8.2 tidyselect_1.2.1
#> [37] digest_0.6.39 future_1.69.0 purrr_1.2.1
#> [40] listenv_0.10.0 labeling_0.4.3 splines_4.5.2
#> [43] fastmap_1.2.0 grid_4.5.2 cli_3.6.5
#> [46] magrittr_2.0.4 patchwork_1.3.2 utf8_1.2.6
#> [49] survival_3.8-3 future.apply_1.20.1 withr_3.0.2
#> [52] scales_1.4.0 lubridate_1.9.4 timechange_0.4.0
#> [55] rmarkdown_2.30 globals_0.19.0 nnet_7.3-20
#> [58] timeDate_4052.112 ragg_1.5.0 evaluate_1.0.5
#> [61] knitr_1.51 hardhat_1.4.2 rlang_1.1.7
#> [64] Rcpp_1.1.1 glue_1.8.0 ipred_0.9-15
#> [67] jsonlite_2.0.0 R6_2.6.1 systemfonts_1.3.1
#> [70] fs_1.6.6