Overview
After processing SEC data with measure.sec, you’ll need to export results for reports, further analysis, or regulatory submissions. This guide covers:
- Creating summary tables with MW averages
- Extracting slice-by-slice data for detailed analysis
- Comparing multiple samples side-by-side
- Generating automated reports
- Exporting to Excel, CSV, and other formats
Quick Reference: Export Functions
| Function | Purpose | Output |
|---|---|---|
measure_sec_summary_table() |
Per-sample MW averages | Tibble with Mn, Mw, Mz, dispersity |
measure_sec_slice_table() |
Point-by-point data | Long or wide format tibble |
measure_sec_compare() |
Multi-sample comparison | Summary, differences, optional plot |
measure_sec_report() |
Automated reports | HTML, PDF, or Word document |
Setup
library(measure)
#> Loading required package: recipes
#> Loading required package: dplyr
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
#>
#> Attaching package: 'recipes'
#> The following object is masked from 'package:stats':
#>
#> step
library(measure.sec)
library(recipes)
library(dplyr)
library(ggplot2)Creating Processed Data for Export
In a real workflow, you’d process SEC data through calibration and MW averaging steps (see the Getting Started vignette). Here, we’ll create representative processed data to demonstrate the export functions.
# Simulate processed SEC results with realistic MW values
# In practice, these columns come from step_sec_mw_averages()
processed <- tibble::tibble(
sample_id = c("PS-50K", "PS-100K", "PS-200K"),
polymer_type = "polystyrene",
# MW averages (typical for narrow PS standards)
mw_mn = c(48500, 97200, 195000),
mw_mw = c(50200, 101000, 203000),
mw_mz = c(52100, 105000, 211000),
mw_dispersity = c(1.035, 1.039, 1.041),
# Additional metadata
injection_volume = 100, # µL
concentration = 2.0 # mg/mL
)
processed
#> # A tibble: 3 × 8
#> sample_id polymer_type mw_mn mw_mw mw_mz mw_dispersity injection_volume
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 PS-50K polystyrene 48500 50200 52100 1.03 100
#> 2 PS-100K polystyrene 97200 101000 105000 1.04 100
#> 3 PS-200K polystyrene 195000 203000 211000 1.04 100
#> # ℹ 1 more variable: concentration <dbl>For multi-sample comparisons, let’s create data for different polymer types:
# Three batches with slight variations (typical QC scenario)
batch1 <- tibble::tibble(
sample_id = "Batch-001",
mw_mn = 48200, mw_mw = 50100, mw_mz = 52000, mw_dispersity = 1.039
)
batch2 <- tibble::tibble(
sample_id = "Batch-002",
mw_mn = 49100, mw_mw = 51200, mw_mz = 53100, mw_dispersity = 1.043
)
batch3 <- tibble::tibble(
sample_id = "Batch-003",
mw_mn = 47800, mw_mw = 49800, mw_mz = 51900, mw_dispersity = 1.042
)Summary Tables
Basic Summary Table
measure_sec_summary_table() creates a one-row-per-sample
table with key metrics:
# Create summary table - automatically finds MW columns
summary_tbl <- measure_sec_summary_table(
processed,
sample_id = "sample_id"
)
print(summary_tbl)
#> SEC Analysis Summary
#> ============================================================
#> Samples: 3
#>
#> # A tibble: 3 × 5
#> sample_id mw_mn mw_mw mw_mz mw_dispersity
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 PS-50K 48500 50200 52100 1.03
#> 2 PS-100K 97200 101000 105000 1.04
#> 3 PS-200K 195000 203000 211000 1.04Including Additional Columns
You can include any numeric column from your data:
# Include method metadata alongside MW results
summary_with_meta <- measure_sec_summary_table(
processed,
sample_id = "sample_id",
additional_cols = c("injection_volume", "concentration")
)
print(summary_with_meta)
#> SEC Analysis Summary
#> ============================================================
#> Samples: 3
#>
#> # A tibble: 3 × 7
#> sample_id mw_mn mw_mw mw_mz mw_dispersity injection_volume concentration
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 PS-50K 48500 50200 52100 1.03 100 2
#> 2 PS-100K 97200 101000 105000 1.04 100 2
#> 3 PS-200K 195000 203000 211000 1.04 100 2Controlling Decimal Places
For regulatory submissions that require specific precision:
# Higher precision for documentation
summary_precise <- measure_sec_summary_table(
processed,
sample_id = "sample_id",
digits = 0 # Whole numbers for MW
)
print(summary_precise)
#> SEC Analysis Summary
#> ============================================================
#> Samples: 3
#>
#> # A tibble: 3 × 5
#> sample_id mw_mn mw_mw mw_mz mw_dispersity
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 PS-50K 48500 50200 52100 1
#> 2 PS-100K 97200 101000 105000 1
#> 3 PS-200K 195000 203000 211000 1Slice Tables: Point-by-Point Data
Understanding Slice Data
SEC analysis works on a point-by-point basis across the chromatogram.
Each “slice” represents one data point with its elution time and
corresponding signal values. Use measure_sec_slice_table()
to extract this detailed data from recipes that produce
measure_list columns.
Creating Slice Data
First, let’s process actual chromatogram data to demonstrate slice extraction:
# Load and process actual chromatogram data
data(sec_triple_detect)
# Process a single sample
ps_sample <- sec_triple_detect |>
filter(sample_id == "PS-100K")
rec <- recipe(
ri_signal + elution_time ~ sample_id,
data = ps_sample
) |>
update_role(sample_id, new_role = "id") |>
step_measure_input_long(
ri_signal,
location = vars(elution_time),
col_name = "ri"
) |>
step_sec_baseline(measures = "ri")
chromatogram_data <- prep(rec) |> bake(new_data = NULL)Long Format (Default)
Long format is best for plotting with ggplot2:
# Extract slice data in long format
slices_long <- measure_sec_slice_table(
chromatogram_data,
measures = "ri",
sample_id = "sample_id"
)
# View structure - one row per data point
head(slices_long, 10)
#> # A tibble: 10 × 5
#> sample_id slice location measure value
#> <chr> <int> <dbl> <chr> <dbl>
#> 1 PS-100K 1 5 ri 0.00132
#> 2 PS-100K 2 5.01 ri 0.00189
#> 3 PS-100K 3 5.02 ri 0.00156
#> 4 PS-100K 4 5.03 ri 0.00123
#> 5 PS-100K 5 5.04 ri 0.00157
#> 6 PS-100K 6 5.05 ri 0.00116
#> 7 PS-100K 7 5.06 ri 0.00188
#> 8 PS-100K 8 5.07 ri 0.00110
#> 9 PS-100K 9 5.08 ri 0.00135
#> 10 PS-100K 10 5.09 ri 0.00126
# Plot using the slice table
ggplot(slices_long, aes(x = location, y = value)) +
geom_line(color = "#2E86AB") +
labs(
x = "Elution Time (min)",
y = "RI Signal (baseline corrected)",
title = "Chromatogram from Slice Data"
) +
theme_minimal()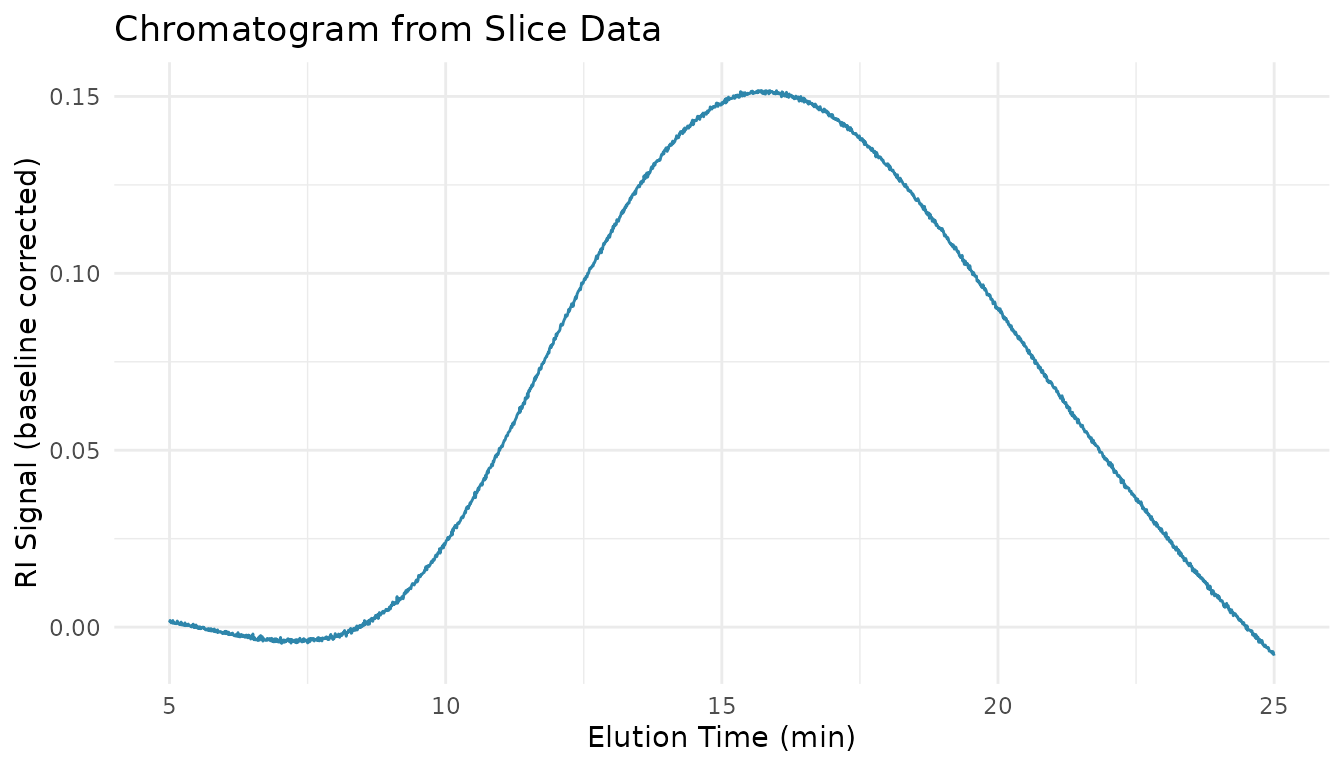
Wide Format
Wide format is better for spreadsheet export or correlating multiple measures:
# Wide format puts each measure in its own column
slices_wide <- measure_sec_slice_table(
chromatogram_data,
measures = "ri",
sample_id = "sample_id",
pivot = TRUE
)
head(slices_wide)
#> # A tibble: 6 × 4
#> sample_id slice location ri
#> <chr> <int> <dbl> <dbl>
#> 1 PS-100K 1 5 0.00132
#> 2 PS-100K 2 5.01 0.00189
#> 3 PS-100K 3 5.02 0.00156
#> 4 PS-100K 4 5.03 0.00123
#> 5 PS-100K 5 5.04 0.00157
#> 6 PS-100K 6 5.05 0.00116Exporting Slice Data to CSV
# Export for external analysis
write.csv(slices_long, "sec_slice_data.csv", row.names = FALSE)
# Or use readr for consistent formatting
readr::write_csv(slices_long, "sec_slice_data.csv")Comparing Multiple Samples
Basic Comparison
measure_sec_compare() provides side-by-side comparison
with differences from a reference:
# Compare the three batches we created earlier
comparison <- measure_sec_compare(
batch1, batch2, batch3,
samples = c("Batch 001", "Batch 002", "Batch 003"),
metrics = "mw_averages",
plot = FALSE
)
print(comparison)
#> SEC Multi-Sample Comparison
#> ============================================================
#> Samples: 3
#> Reference: Batch 001
#>
#> Summary:
#> # A tibble: 3 × 5
#> sample mw_mn mw_mw mw_mz mw_dispersity
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 Batch 001 48200 50100 52000 1.04
#> 2 Batch 002 49100 51200 53100 1.04
#> 3 Batch 003 47800 49800 51900 1.04
#>
#> Differences from reference:
#> # A tibble: 3 × 9
#> sample mw_mn_diff mw_mn_pct mw_mw_diff mw_mw_pct mw_mz_diff mw_mz_pct
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 Batch 001 0 0 0 0 0 0
#> 2 Batch 002 900 1.9 1100 2.2 1100 2.1
#> 3 Batch 003 -400 -0.8 -300 -0.6 -100 -0.2
#> # ℹ 2 more variables: mw_dispersity_diff <dbl>, mw_dispersity_pct <dbl>Understanding the Comparison Output
The comparison object contains:
-
$summary: All samples with their metrics -
$differences: Absolute and percent differences from reference -
$plot: MWD overlay (if requested and available) -
$reference: Which sample is the reference
# Access individual components
comparison$summary # Metrics for all samples
comparison$differences # Differences from reference
comparison$reference # Reference sample nameSetting a Different Reference
By default, the first sample is the reference. You can change this:
# Use PS 500K as reference
comparison_ref <- measure_sec_compare(
ps100k, ps500k, pmma,
samples = c("PS 100K", "PS 500K", "PMMA High"),
reference = "PS 500K"
)Batch-to-Batch Comparison
A common use case is comparing production batches to a reference lot:
# Compare production batches
batch_comparison <- measure_sec_compare(
reference_lot,
batch_001,
batch_002,
batch_003,
samples = c("Reference", "Batch 001", "Batch 002", "Batch 003"),
reference = "Reference"
)
# Check for significant deviations
batch_comparison$differences |>
filter(abs(mw_mw_pct) > 5) # Flag batches > 5% differentAutomated Reports
Available Templates
# See available report templates
list_sec_templates()
#> # A tibble: 3 × 3
#> template description formats
#> <chr> <chr> <chr>
#> 1 standard Summary table, chromatogram, and MWD plot html, pdf, docx
#> 2 detailed All plots, multi-detector view, optional slice data html, pdf, docx
#> 3 qc System suitability with pass/fail metrics html, pdf, docxStandard Report
The standard template includes: - Summary table with MW averages - Chromatogram overlay - Molecular weight distribution plot
# Generate HTML report
measure_sec_report(
processed,
template = "standard",
output_format = "html",
title = "Polymer SEC Analysis",
author = "Lab Analyst"
)Detailed Report
For more comprehensive documentation:
measure_sec_report(
processed,
template = "detailed",
output_format = "pdf",
title = "Comprehensive SEC Report",
sample_id = "sample_id",
include_slice_table = TRUE # Append raw data
)QC Report
For system suitability testing:
measure_sec_report(
sst_data,
template = "qc",
output_format = "html",
specs = list(
plate_count_min = 10000,
asymmetry_min = 0.8,
asymmetry_max = 1.5,
resolution_min = 1.5
)
)Saving Reports to Specific Locations
# Save to a specific path
measure_sec_report(
processed,
template = "standard",
output_file = "reports/polymer_analysis_2024-01-15.html",
open = FALSE # Don't open automatically
)Integration with Other Tools
Export for GraphPad Prism
Prism prefers wide format with specific column arrangements:
# Create Prism-friendly format
prism_data <- slices_long |>
select(location, sample_id, value) |>
tidyr::pivot_wider(names_from = sample_id, values_from = value)
write.csv(prism_data, "sec_for_prism.csv", row.names = FALSE)Best Practices
Traceability
Always include metadata for regulatory compliance:
# Add audit trail information
summary_with_audit <- summary_tbl |>
mutate(
analysis_date = Sys.Date(),
analyst = "JW",
instrument = "Agilent 1260",
column = "PLgel Mixed-C",
software_version = packageVersion("measure.sec")
)File Naming Conventions
Use descriptive, consistent file names:
# Good: includes key information
"sec_summary_batch123_2024-01-15.xlsx"
"mwd_comparison_stability_t12m.pdf"
# Avoid: ambiguous names
"results.xlsx"
"output.csv"Version Control for Reports
Save report parameters for reproducibility:
# Save analysis parameters alongside results
analysis_params <- list(
date = Sys.time(),
calibration = "ps_2024-01.rds",
baseline_method = "linear",
integration_limits = c(8, 18),
package_version = packageVersion("measure.sec")
)
saveRDS(analysis_params, "sec_analysis_params.rds")Troubleshooting
Missing Columns in Summary
If MW columns are missing from your summary table, check that you’ve run the calibration and MW averaging steps:
# Ensure you have MW data
names(processed) # Should include Mn, Mw, Mz, or mw_mn, mw_mw, mw_mzEmpty Slice Tables
If measure_sec_slice_table() returns no data:
# Check that measure columns exist
find_measure_cols <- function(data) {
names(data)[vapply(data, inherits, logical(1), "measure_list")]
}
find_measure_cols(processed)Report Generation Fails
If measure_sec_report() fails:
- Verify Quarto is installed:
quarto::quarto_version() - For PDF output, ensure LaTeX is installed
- Check data has required columns
See Also
- Getting Started - Basic SEC workflow and concepts
- Calibration Management - Save and reuse calibrations
- System Suitability - QC metrics and column performance
- Comprehensive SEC Analysis - Complete reference with all functions
Session Info
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.1 measure.sec_0.0.0.9000 measure_0.0.1.9002
#> [4] recipes_1.3.1 dplyr_1.1.4
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.56 bslib_0.10.0
#> [4] lattice_0.22-7 vctrs_0.7.1 tools_4.5.2
#> [7] generics_0.1.4 parallel_4.5.2 tibble_3.3.1
#> [10] pkgconfig_2.0.3 Matrix_1.7-4 data.table_1.18.2.1
#> [13] RColorBrewer_1.1-3 S7_0.2.1 desc_1.4.3
#> [16] lifecycle_1.0.5 compiler_4.5.2 farver_2.1.2
#> [19] textshaping_1.0.4 codetools_0.2-20 htmltools_0.5.9
#> [22] class_7.3-23 sass_0.4.10 yaml_2.3.12
#> [25] prodlim_2025.04.28 tidyr_1.3.2 pillar_1.11.1
#> [28] pkgdown_2.2.0 jquerylib_0.1.4 MASS_7.3-65
#> [31] cachem_1.1.0 gower_1.0.2 rpart_4.1.24
#> [34] parallelly_1.46.1 lava_1.8.2 tidyselect_1.2.1
#> [37] digest_0.6.39 future_1.69.0 purrr_1.2.1
#> [40] listenv_0.10.0 labeling_0.4.3 splines_4.5.2
#> [43] fastmap_1.2.0 grid_4.5.2 cli_3.6.5
#> [46] magrittr_2.0.4 utf8_1.2.6 survival_3.8-3
#> [49] future.apply_1.20.1 withr_3.0.2 scales_1.4.0
#> [52] lubridate_1.9.4 timechange_0.4.0 rmarkdown_2.30
#> [55] globals_0.19.0 nnet_7.3-20 timeDate_4052.112
#> [58] ragg_1.5.0 evaluate_1.0.5 knitr_1.51
#> [61] hardhat_1.4.2 rlang_1.1.7 Rcpp_1.1.1
#> [64] glue_1.8.0 ipred_0.9-15 jsonlite_2.0.0
#> [67] R6_2.6.1 systemfonts_1.3.1 fs_1.6.6