Single-Angle Light Scattering: LALS and RALS
Source:vignettes/lals-rals-detection.Rmd
lals-rals-detection.RmdOverview
Single-angle light scattering detectors (LALS and RALS) provide a simpler, cost-effective alternative to multi-angle detection (MALS) for determining absolute molecular weight. While they don’t provide radius of gyration (Rg) information, they can be excellent choices for routine analysis of smaller molecules.
This vignette covers:
- LALS (Low-Angle Light Scattering) principles and usage
- RALS (Right-Angle Light Scattering) principles and usage
- When to choose each technique
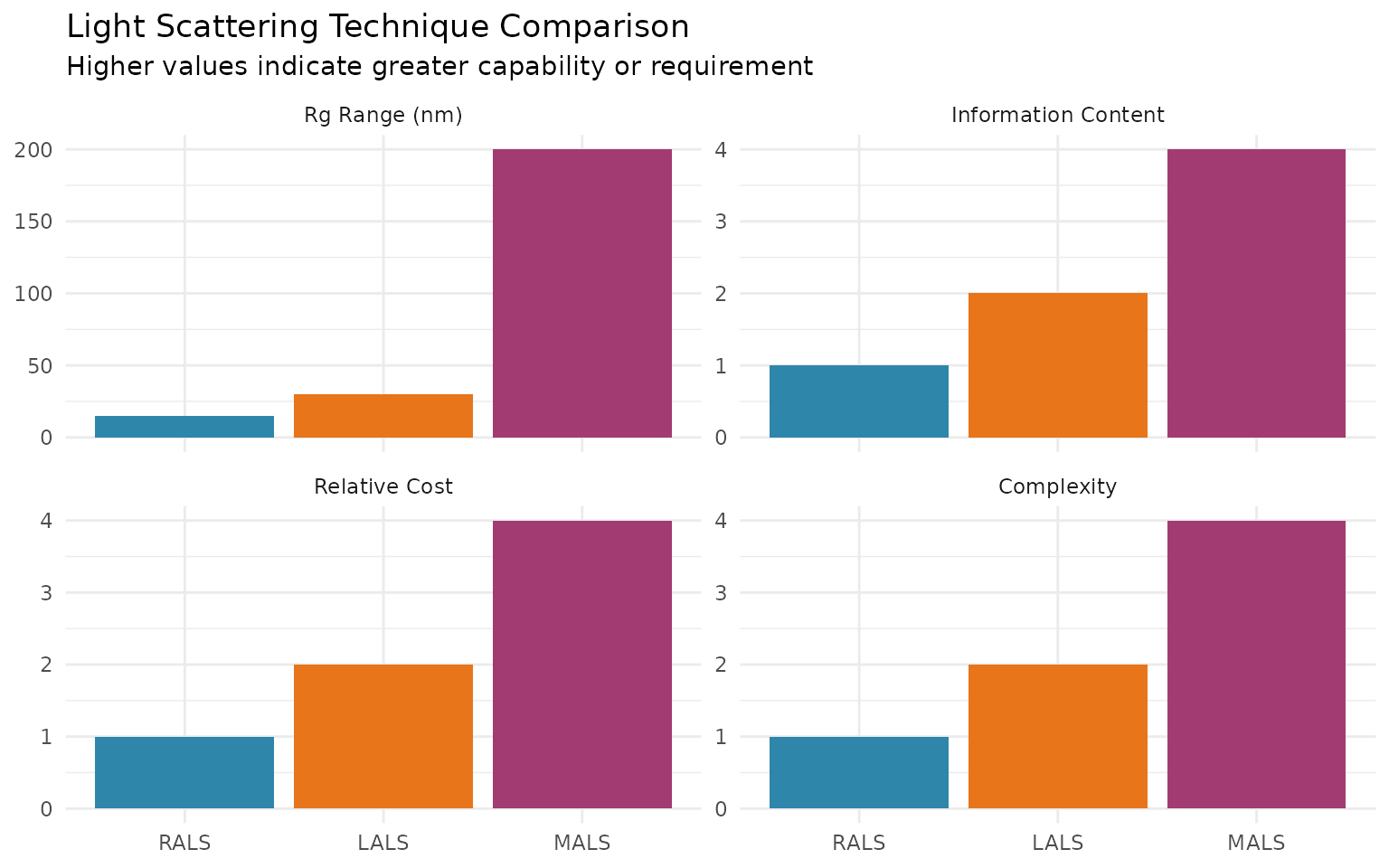
- Comparison with MALS
Setup
library(measure)
#> Loading required package: recipes
#> Loading required package: dplyr
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
#>
#> Attaching package: 'recipes'
#> The following object is masked from 'package:stats':
#>
#> step
library(measure.sec)
library(recipes)
library(dplyr)
library(ggplot2)The Angular Dependence Problem
Light scattering from polymer molecules is angle-dependent. For larger molecules, the scattered intensity varies with detection angle due to intramolecular interference. This is described by the particle scattering function P(θ):
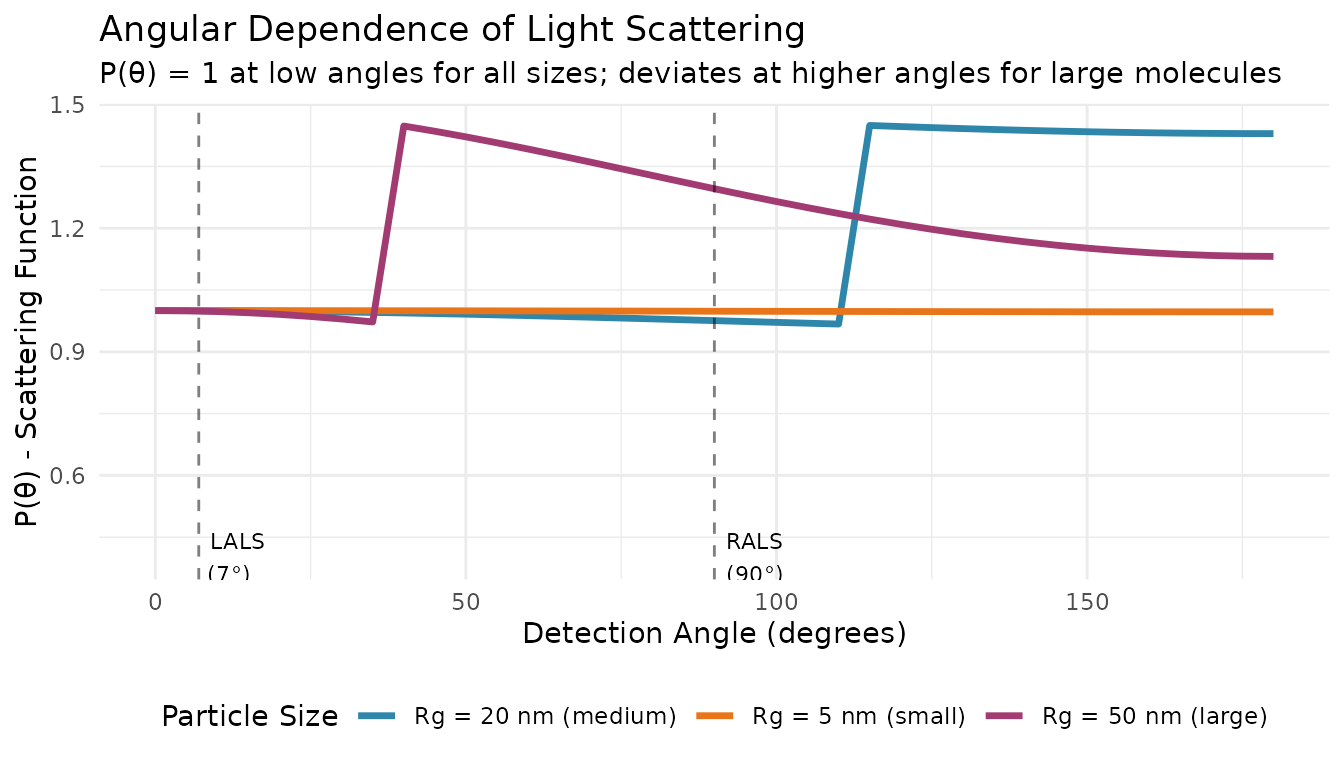
Key insight: At low angles (approaching 0°), P(θ) → 1 regardless of particle size. This is why LALS can measure MW without knowing Rg.
LALS: Low-Angle Light Scattering
Principle
LALS detectors measure scattering at a low angle (typically 7-15°). At these angles, the particle scattering function P(θ) ≈ 1 for most polymer sizes, eliminating the need for angular extrapolation.
The simplified equation becomes:
Where: - R(θ) = Rayleigh ratio (scattered intensity) - K = optical constant - c = concentration
When to Use LALS
| Advantage | Limitation |
|---|---|
| No angular extrapolation needed | No Rg information |
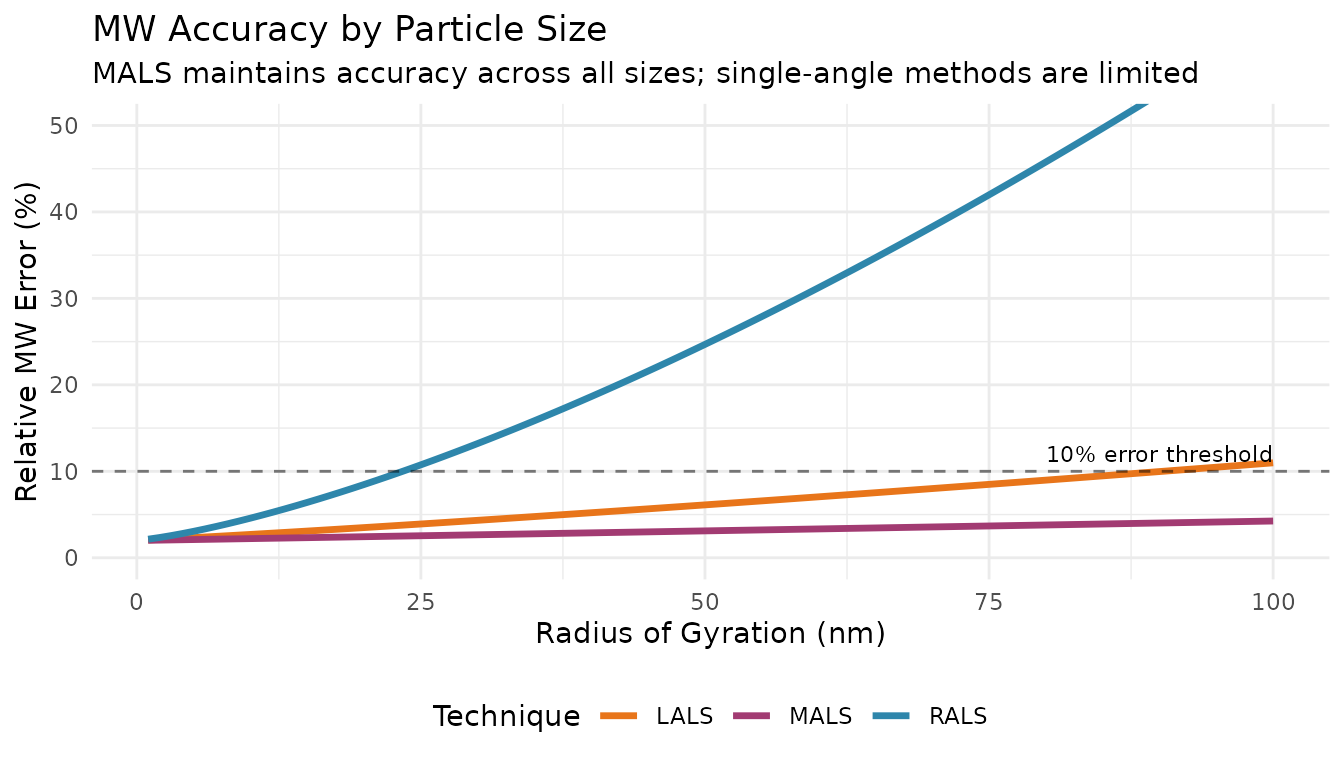
| Simple, single-angle measurement | Less accurate for very large molecules |
| Faster data acquisition | Requires calibration constant |
| Lower cost than MALS | More sensitive to dust/aggregates |
Best for: - Proteins and peptides (Rg < 10 nm) - Small synthetic polymers - Routine screening applications - When Rg is not needed
LALS Workflow
# LALS workflow example
# Note: Requires LALS detector data
# Simulate sample data structure
lals_samples <- tibble(
sample_id = "Protein-A",
ri_signal = list(c(0.1, 0.5, 1.0, 0.5, 0.1)),
lals_signal = list(c(0.05, 0.25, 0.5, 0.25, 0.05)),
elution_time = list(c(8, 9, 10, 11, 12)),
dn_dc = 0.185
)
rec_lals <- recipe(
ri_signal + lals_signal + elution_time + dn_dc ~ sample_id,
data = lals_samples
) |>
update_role(sample_id, new_role = "id") |>
# Convert signals to measure format
step_measure_input_long(ri_signal, location = vars(elution_time), col_name = "ri") |>
step_measure_input_long(lals_signal, location = vars(elution_time), col_name = "lals") |>
# Baseline correction
step_sec_baseline(measures = c("ri", "lals")) |>
# Process RI for concentration
step_sec_ri(measures = "ri", dn_dc_column = "dn_dc") |>
step_sec_concentration(
measures = "ri",
detector = "ri",
injection_volume = 100,
sample_concentration = 2.0
) |>
# Process LALS for MW
step_sec_lals(
measures = "lals",
concentration_col = "ri",
angle = 7, # Detection angle
laser_wavelength = 670, # nm
dn_dc = 0.185, # mL/g
solvent_ri = 1.333, # Water
calibration_constant = 1.5e-5 # From instrument calibration
)
prepped_lals <- prep(rec_lals)
result_lals <- bake(prepped_lals, new_data = NULL)RALS: Right-Angle Light Scattering
Principle
RALS detectors measure scattering at 90°. While this angle shows more angular dependence than low angles, it offers practical advantages: less sensitivity to stray light and easier optical design.
For small molecules where Rg << λ/20 (roughly Rg < 15 nm at 670 nm), the angular dependence is minimal and RALS provides reasonable MW estimates.
When to Use RALS
| Advantage | Limitation |
|---|---|
| Less sensitive to dust/stray light | Underestimates MW for large molecules |
| Common, cost-effective detector | No Rg information |
| Good for small molecules | Angular correction needed for large MW |
| Often included in GPC systems | Less accurate than LALS for medium sizes |
Best for: - Small molecules (Rg < 15 nm) - Quality control screening - Cost-sensitive applications - When combined with other detectors
RALS Workflow
# RALS workflow example
# Note: Requires RALS detector data
# Simulate sample data structure
rals_samples <- tibble(
sample_id = "Polymer-X",
ri_signal = list(c(0.1, 0.5, 1.0, 0.5, 0.1)),
rals_signal = list(c(0.04, 0.20, 0.40, 0.20, 0.04)),
elution_time = list(c(8, 9, 10, 11, 12)),
dn_dc = 0.185
)
rec_rals <- recipe(
ri_signal + rals_signal + elution_time + dn_dc ~ sample_id,
data = rals_samples
) |>
update_role(sample_id, new_role = "id") |>
# Convert signals to measure format
step_measure_input_long(ri_signal, location = vars(elution_time), col_name = "ri") |>
step_measure_input_long(rals_signal, location = vars(elution_time), col_name = "rals") |>
# Baseline correction
step_sec_baseline(measures = c("ri", "rals")) |>
# Process RI for concentration
step_sec_ri(measures = "ri", dn_dc_column = "dn_dc") |>
step_sec_concentration(
measures = "ri",
detector = "ri",
injection_volume = 100,
sample_concentration = 2.0
) |>
# Process RALS for MW
step_sec_rals(
measures = "rals",
concentration_col = "ri",
angle = 90, # Right angle
laser_wavelength = 670, # nm
dn_dc = 0.185, # mL/g
solvent_ri = 1.333, # Water
calibration_constant = 1.2e-5 # From instrument calibration
)
prepped_rals <- prep(rec_rals)
result_rals <- bake(prepped_rals, new_data = NULL)Choosing the Right Technique
Combining LALS/RALS with MALS
Some instruments include both low-angle and right-angle detectors alongside MALS. This provides:
- Redundancy: Cross-check MW values
- Extended range: LALS for low angles, MALS for extrapolation
- Sensitivity optimization: Use optimal detector for each MW range
# Combined LALS + RALS + MALS workflow
rec_combined <- recipe(
ri_signal + lals_signal + rals_signal + mals_signal + elution_time + dn_dc ~ sample_id,
data = multi_detector_samples
) |>
update_role(sample_id, new_role = "id") |>
# Input all detectors
step_measure_input_long(ri_signal, location = vars(elution_time), col_name = "ri") |>
step_measure_input_long(lals_signal, location = vars(elution_time), col_name = "lals") |>
step_measure_input_long(rals_signal, location = vars(elution_time), col_name = "rals") |>
step_measure_input_long(mals_signal, location = vars(elution_time), col_name = "mals") |>
# Baseline and concentration
step_sec_baseline(measures = c("ri", "lals", "rals", "mals")) |>
step_sec_ri(measures = "ri", dn_dc_column = "dn_dc") |>
step_sec_concentration(
measures = "ri",
detector = "ri",
injection_volume = 100,
sample_concentration = 2.0
) |>
# Process each LS detector
step_sec_lals(measures = "lals", concentration_col = "ri", dn_dc = 0.185) |>
step_sec_rals(measures = "rals", concentration_col = "ri", dn_dc = 0.185) |>
step_sec_mals(mals_col = "mals", dn_dc_column = "dn_dc")Calibration Constants
Both step_sec_lals() and step_sec_rals()
require a calibration_constant for absolute MW values.
Without it, results are in relative units.
Determining Calibration Constants
- Use a well-characterized standard (e.g., BSA, narrow PS standard)
- Run the standard under your experimental conditions
- Calculate the constant that yields the known MW
# Example: Calibrating with BSA (MW = 66,430 Da)
# 1. Measure BSA with known concentration
# 2. Calculate: cal_constant = (known_MW * K * c) / raw_signal
bsa_mw <- 66430 # Da
bsa_conc <- 1.0 # mg/mL
dn_dc_bsa <- 0.185
solvent_ri <- 1.333
wavelength <- 670
# Calculate K
K <- 4 * pi^2 * solvent_ri^2 * dn_dc_bsa^2 / (6.022e23 * (wavelength * 1e-7)^4)
# If raw_signal at peak = 0.5, then:
# cal_constant = bsa_mw * K * bsa_conc / raw_signalTroubleshooting
See Also
- MALS Detection - Multi-angle detection for Rg and large molecules
- Triple Detection - Combined RI + Viscometer + LS workflows
- Getting Started - Basic SEC workflow and concepts
- Calibration Management - Save and reuse calibrations
Session Info
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.1 measure.sec_0.0.0.9000 measure_0.0.1.9002
#> [4] recipes_1.3.1 dplyr_1.1.4
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.56 bslib_0.10.0
#> [4] lattice_0.22-7 vctrs_0.7.1 tools_4.5.2
#> [7] generics_0.1.4 parallel_4.5.2 tibble_3.3.1
#> [10] pkgconfig_2.0.3 Matrix_1.7-4 data.table_1.18.2.1
#> [13] RColorBrewer_1.1-3 S7_0.2.1 desc_1.4.3
#> [16] lifecycle_1.0.5 compiler_4.5.2 farver_2.1.2
#> [19] textshaping_1.0.4 codetools_0.2-20 htmltools_0.5.9
#> [22] class_7.3-23 sass_0.4.10 yaml_2.3.12
#> [25] prodlim_2025.04.28 tidyr_1.3.2 pillar_1.11.1
#> [28] pkgdown_2.2.0 jquerylib_0.1.4 MASS_7.3-65
#> [31] cachem_1.1.0 gower_1.0.2 rpart_4.1.24
#> [34] parallelly_1.46.1 lava_1.8.2 tidyselect_1.2.1
#> [37] digest_0.6.39 future_1.69.0 purrr_1.2.1
#> [40] listenv_0.10.0 labeling_0.4.3 splines_4.5.2
#> [43] fastmap_1.2.0 grid_4.5.2 cli_3.6.5
#> [46] magrittr_2.0.4 survival_3.8-3 future.apply_1.20.1
#> [49] withr_3.0.2 scales_1.4.0 lubridate_1.9.4
#> [52] timechange_0.4.0 rmarkdown_2.30 globals_0.19.0
#> [55] nnet_7.3-20 timeDate_4052.112 ragg_1.5.0
#> [58] evaluate_1.0.5 knitr_1.51 hardhat_1.4.2
#> [61] rlang_1.1.7 Rcpp_1.1.1 glue_1.8.0
#> [64] ipred_0.9-15 jsonlite_2.0.0 R6_2.6.1
#> [67] systemfonts_1.3.1 fs_1.6.6