Multi-Angle Light Scattering (MALS) for Absolute Molecular Weight
Source:vignettes/mals-detection.Rmd
mals-detection.RmdOverview
Multi-Angle Light Scattering (MALS) is a powerful technique that provides absolute molecular weight without calibration standards. Unlike conventional calibration (which gives MW relative to polymer standards), MALS measures the actual molecular weight of your sample based on fundamental light scattering physics. This vignette covers:
How MALS measures absolute molecular weight
Key parameters: dn/dc, wavelength, angles
Zimm, Debye, and Berry formalisms
Radius of gyration (Rg) from angular dependence
Practical workflows with
step_sec_mals()
Setup
library(measure)
#> Loading required package: recipes
#> Loading required package: dplyr
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
#>
#> Attaching package: 'recipes'
#> The following object is masked from 'package:stats':
#>
#> step
library(measure.sec)
library(recipes)
library(dplyr)
library(ggplot2)How MALS Works
The Light Scattering Principle
When light passes through a polymer solution, molecules scatter light in all directions. The intensity of scattered light depends on:
- Molecular weight: Larger molecules scatter more light
- Concentration: More molecules = more scattering
- Scattering angle: Angular dependence reveals molecular size
The fundamental relationship is the Rayleigh-Debye equation:
Where:
| Symbol | Meaning |
|---|---|
| K | Optical constant (depends on dn/dc, wavelength, solvent) |
| c | Concentration (g/mL) |
| R(θ) | Excess Rayleigh ratio (scattered intensity) |
| Mw | Weight-average molecular weight |
| P(θ) | Particle scattering function (angular dependence) |
| A₂ | Second virial coefficient |
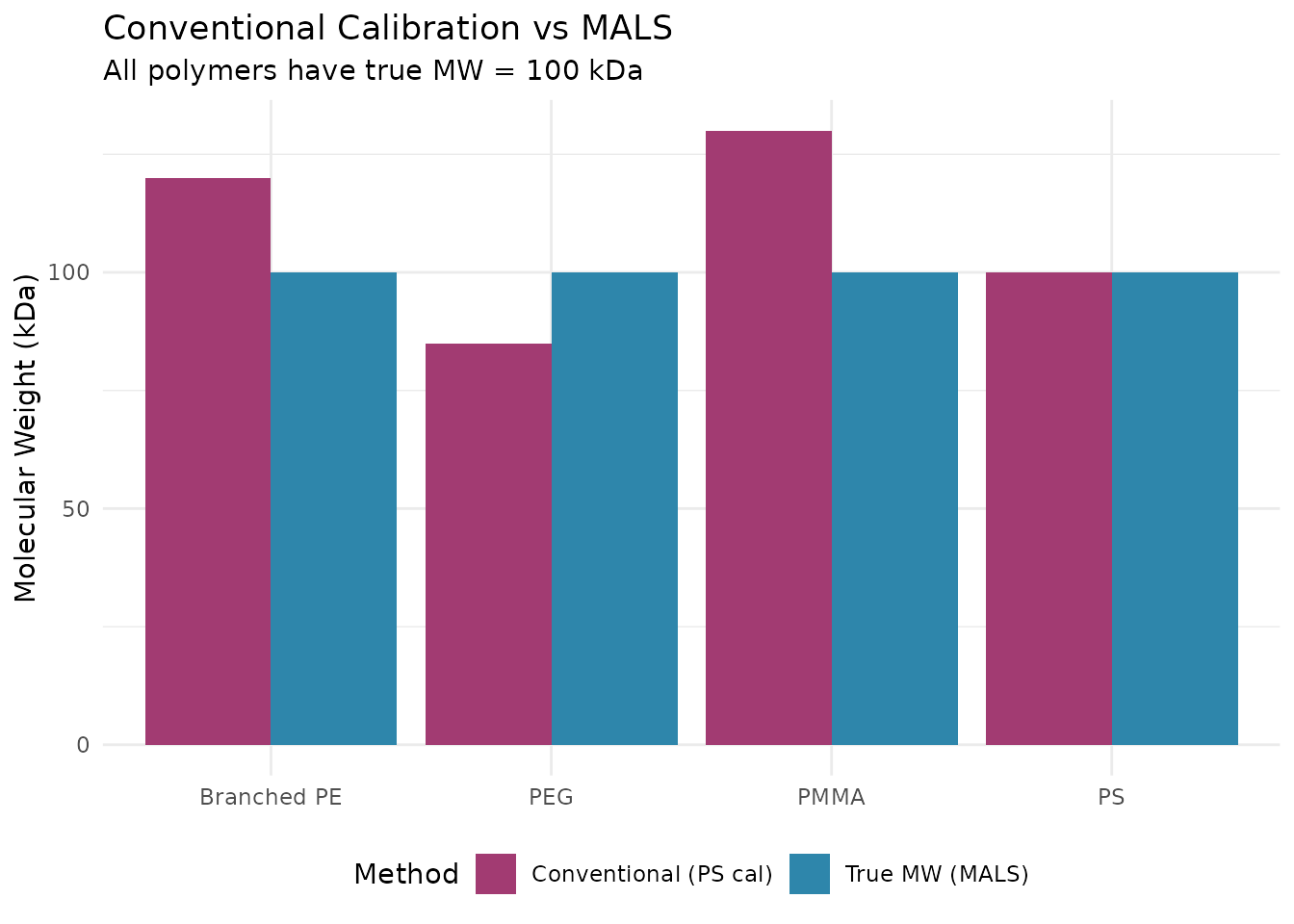
Why Use MALS?
Conventional Calibration Limitations
Conventional (relative) calibration has significant limitations:
| Polymer Type | Conventional Cal | MALS |
|---|---|---|
| Same as standards | Accurate | Accurate |
| Different chemistry | Biased | Accurate |
| Branched | Biased | Accurate |
| Copolymers | Biased | Accurate |
MALS is essential when: - Analyzing polymers different from your calibration standards - Studying branched or star polymers - Characterizing copolymers - Absolute MW is required for regulatory submissions
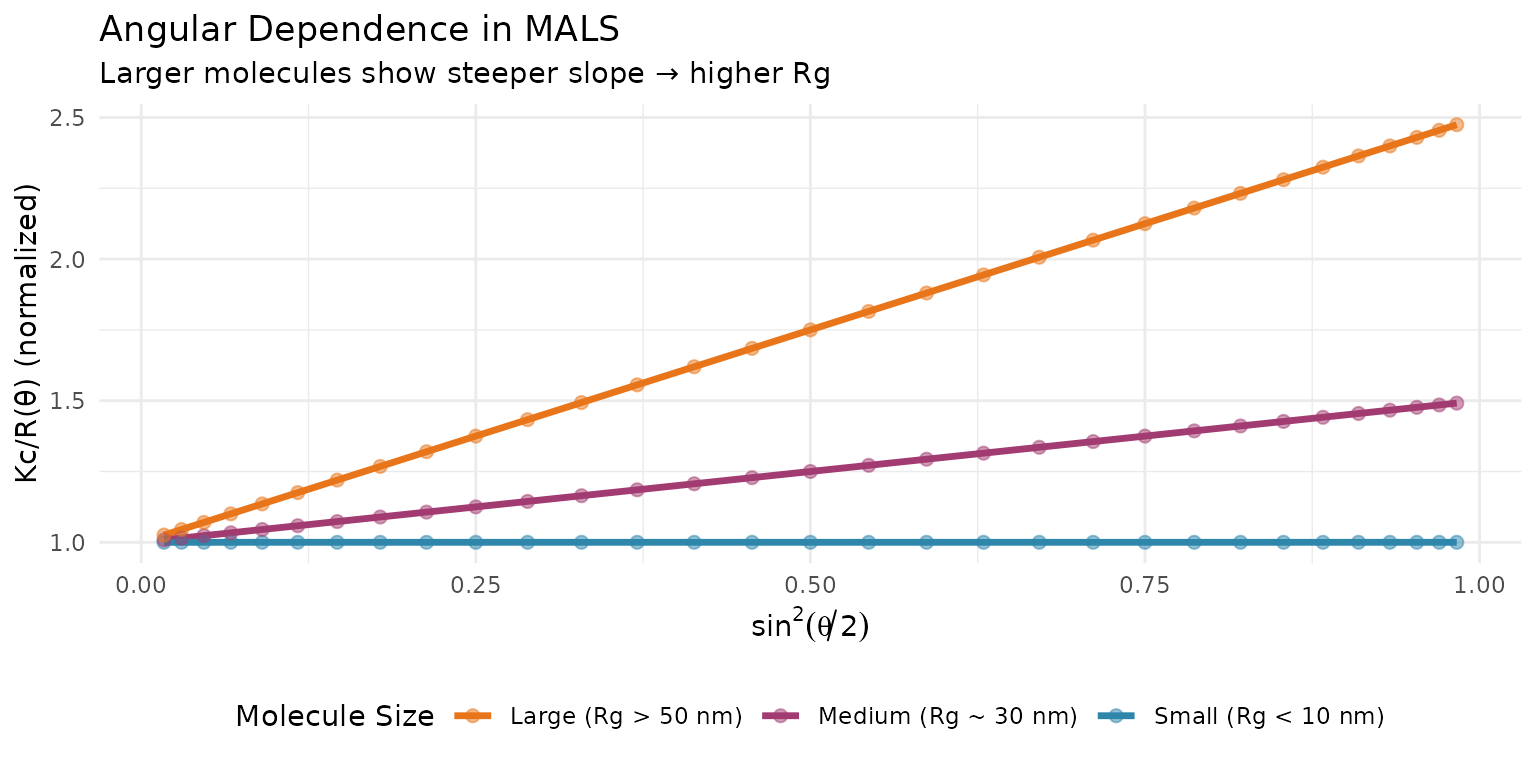
Angular Dependence and Rg
Formalisms for Angular Extrapolation
Zimm, Debye, and Berry
Three common approaches for angular extrapolation:
| Formalism | Plot | Best For |
|---|---|---|
| Zimm | Kc/R vs sin²(θ/2) | Random coils, most polymers |
| Debye | Kc/R vs sin²(θ/2) | Similar to Zimm |
| Berry | √(Kc/R) vs sin²(θ/2) | Very large particles, aggregates |
#> `geom_smooth()` using formula = 'y ~ x'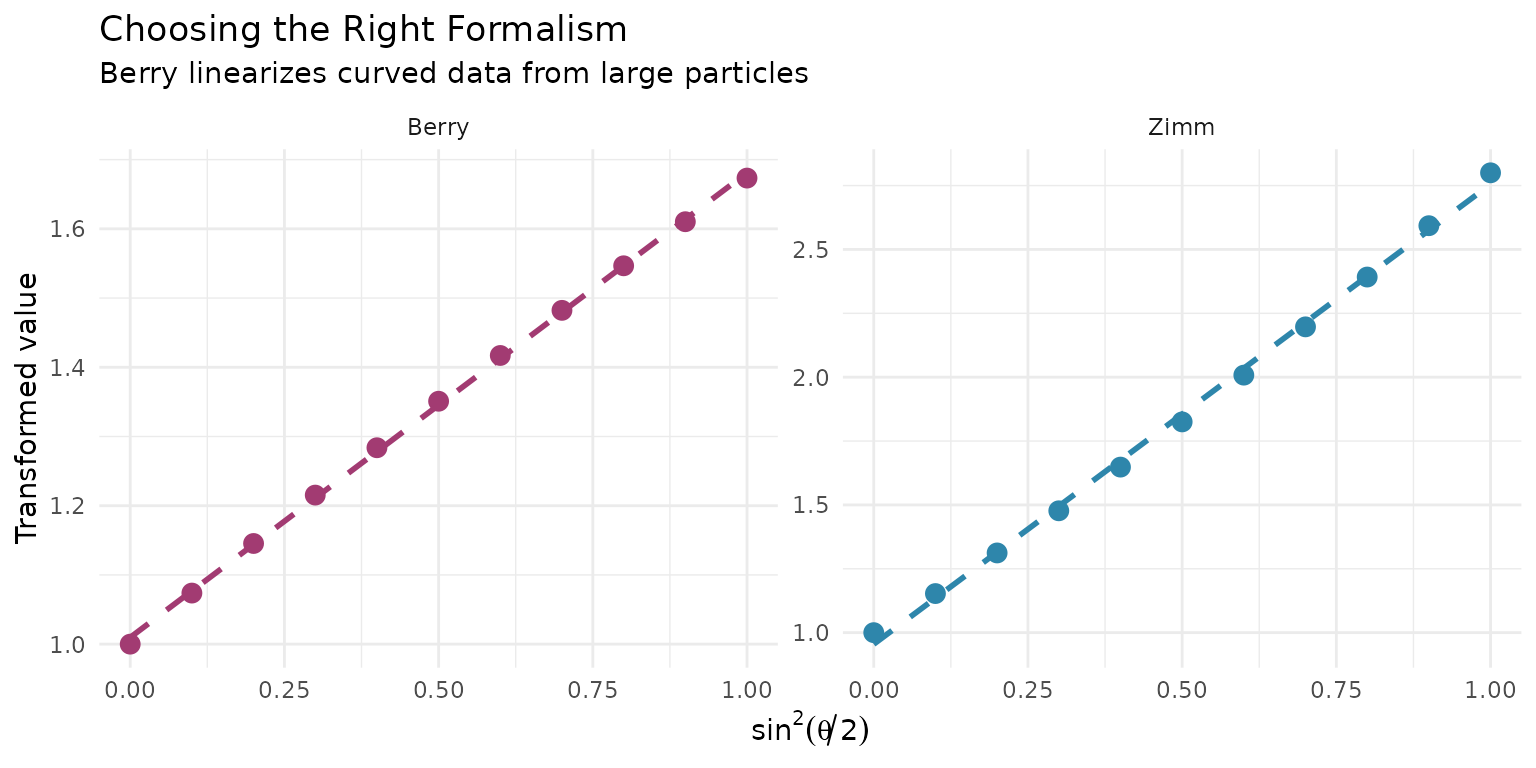
Choosing a formalism:
- Start with Zimm - works for most polymers
- Use Berry if Zimm plot shows significant curvature
- Berry is essential for Rg > 50 nm or aggregates
Example Dataset
data(sec_triple_detect, package = "measure.sec")
# Select samples with MALS data
mals_samples <- sec_triple_detect |>
filter(sample_type == "sample")
glimpse(mals_samples)
#> Rows: 14,007
#> Columns: 11
#> $ sample_id <chr> "PMMA-Low", "PMMA-Low", "PMMA-Low", "PMMA-Low", "PMMA…
#> $ sample_type <chr> "sample", "sample", "sample", "sample", "sample", "sa…
#> $ polymer_type <chr> "pmma", "pmma", "pmma", "pmma", "pmma", "pmma", "pmma…
#> $ elution_time <dbl> 5.00, 5.01, 5.02, 5.03, 5.04, 5.05, 5.06, 5.07, 5.08,…
#> $ ri_signal <dbl> 2.177879e-04, 0.000000e+00, 2.307149e-04, 1.490633e-0…
#> $ uv_signal <dbl> 0.000000e+00, 0.000000e+00, 0.000000e+00, 6.442527e-0…
#> $ mals_signal <dbl> 3.454417e-06, 1.210776e-05, 1.804800e-05, 2.001408e-0…
#> $ known_mw <dbl> 25000, 25000, 25000, 25000, 25000, 25000, 25000, 2500…
#> $ known_dispersity <dbl> 1.8, 1.8, 1.8, 1.8, 1.8, 1.8, 1.8, 1.8, 1.8, 1.8, 1.8…
#> $ dn_dc <dbl> 0.084, 0.084, 0.084, 0.084, 0.084, 0.084, 0.084, 0.08…
#> $ extinction_coef <dbl> 0.1, 0.1, 0.1, 0.1, 0.1, 0.1, 0.1, 0.1, 0.1, 0.1, 0.1…MALS Workflow with measure.sec
Basic MALS Analysis
# Complete MALS workflow for absolute MW
rec_mals <- recipe(
ri_signal + mals_signal + elution_time + dn_dc ~ sample_id,
data = mals_samples
) |>
update_role(sample_id, new_role = "id") |>
# Convert signals to measure format
step_measure_input_long(ri_signal, location = vars(elution_time), col_name = "ri") |>
step_measure_input_long(mals_signal, location = vars(elution_time), col_name = "mals") |>
# Baseline correction
step_sec_baseline(measures = c("ri", "mals")) |>
# Process RI for concentration
step_sec_ri(measures = "ri", dn_dc_column = "dn_dc") |>
# Convert RI to concentration
step_sec_concentration(
measures = "ri",
detector = "ri",
injection_volume = 100, # µL
sample_concentration = 2.0 # mg/mL
) |>
# MALS processing for absolute MW
step_sec_mals(
mals_col = "mals",
dn_dc_column = "dn_dc",
wavelength = 658, # nm (common MALS laser)
solvent_ri = 1.407, # THF
angles = 90 # Single angle for this example
)
prepped_mals <- prep(rec_mals)
result_mals <- bake(prepped_mals, new_data = NULL)
# View results
result_mals |>
select(sample_id, ri, mals, mw_mals) |>
head(3)
#> # A tibble: 3 × 4
#> sample_id ri mals mw_mals
#> <chr> <meas> <meas> <meas>
#> 1 PMMA-Low [2,001 × 2] [2,001 × 2] [2,001 × 2]
#> 2 PMMA-Med [2,001 × 2] [2,001 × 2] [2,001 × 2]
#> 3 PMMA-High [2,001 × 2] [2,001 × 2] [2,001 × 2]Key Parameters
| Parameter | Description | Typical Values |
|---|---|---|
dn_dc |
Refractive index increment | 0.185 (PS/THF), 0.084 (PMMA/THF) |
wavelength |
Laser wavelength (nm) | 658, 690, 785 |
solvent_ri |
Solvent refractive index | 1.333 (water), 1.407 (THF) |
angles |
Detection angle(s) | 90 (single), c(35, 50, 75, 90, 105, 120, 145) (multi) |
formalism |
Extrapolation method | “zimm”, “debye”, “berry” |
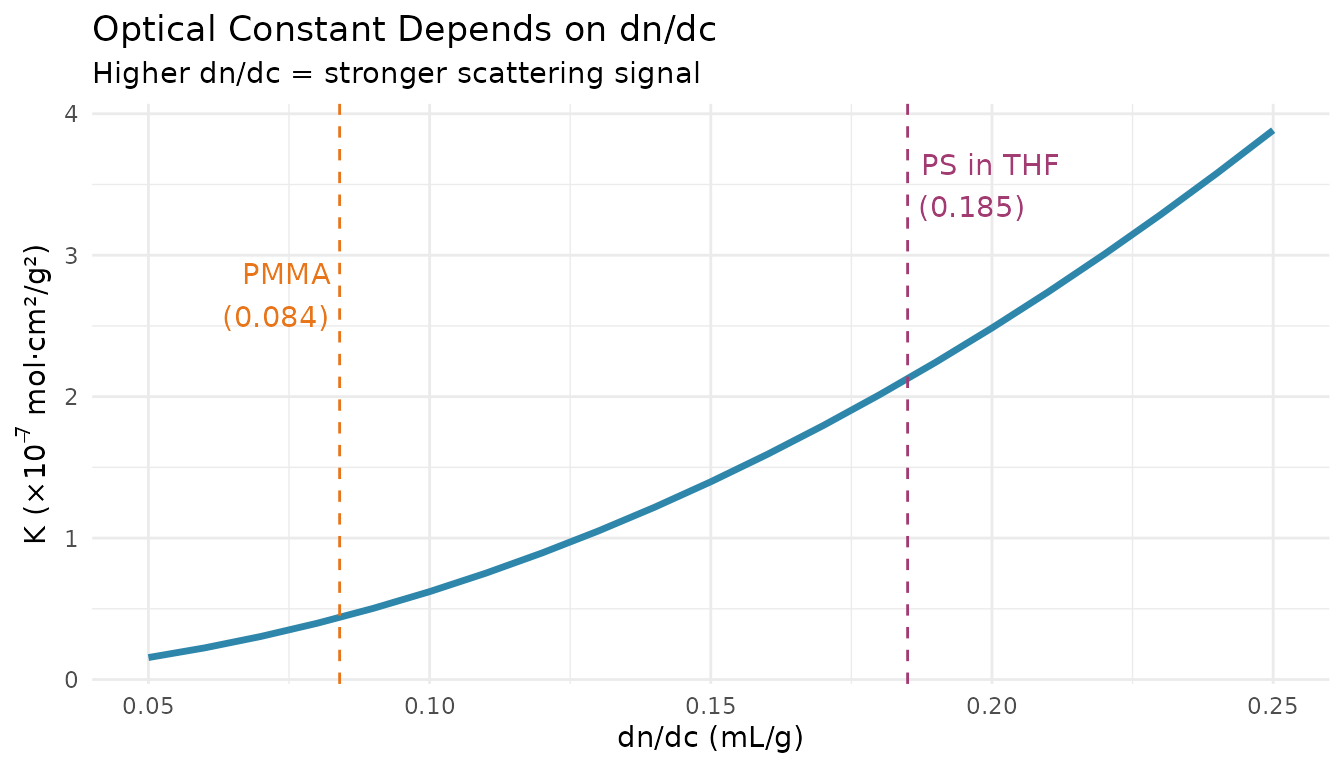
The Critical Role of dn/dc
What is dn/dc?
The refractive index increment (dn/dc) is the change in solution refractive index per unit concentration:
Multi-Angle Detection
Setting Up Multi-Angle Analysis
For Rg determination, provide multiple detection angles:
# Multi-angle MALS for MW and Rg
step_sec_mals(
mals_col = "mals",
dn_dc = 0.185,
wavelength = 658,
solvent_ri = 1.407,
angles = c(35, 50, 75, 90, 105, 120, 145), # Typical MALS detector angles
formalism = "zimm" # Try "berry" for large particles
)Troubleshooting
Common MALS Issues
| Problem | Possible Causes | Solutions |
|---|---|---|
| Noisy MW data | Low concentration, dust | Increase conc, filter samples |
| Negative MW | Baseline issues | Improve baseline, check alignment |
| MW too high | Aggregates, dust | Filter samples, check for aggregation |
| MW too low | Wrong dn/dc | Measure dn/dc accurately |
| Inconsistent Rg | Poor signal, few angles | Use more angles, increase concentration |
When to Use MALS vs Conventional Calibration
| Scenario | Recommendation |
|---|---|
| Routine QC of known polymer | Conventional cal is faster and sufficient |
| Different polymer from standards | Use MALS |
| Branched or complex architecture | Use MALS |
| Absolute MW required | Use MALS |
| Characterizing new materials | Use MALS |
| Regulatory submissions | Use MALS for accuracy |
See Also
- Getting Started - Basic SEC workflow
- LALS and RALS Detection - Simpler light scattering alternatives
- Multi-Detector SEC - RI + Viscometer + LS workflows
- System Suitability - QC metrics for MALS
Session Info
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.3 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.1 measure.sec_0.0.0.9000 measure_0.0.1.9002
#> [4] recipes_1.3.1 dplyr_1.1.4
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.56 bslib_0.10.0
#> [4] lattice_0.22-7 vctrs_0.7.1 tools_4.5.2
#> [7] generics_0.1.4 parallel_4.5.2 tibble_3.3.1
#> [10] pkgconfig_2.0.3 Matrix_1.7-4 data.table_1.18.2.1
#> [13] RColorBrewer_1.1-3 S7_0.2.1 desc_1.4.3
#> [16] lifecycle_1.0.5 compiler_4.5.2 farver_2.1.2
#> [19] textshaping_1.0.4 codetools_0.2-20 htmltools_0.5.9
#> [22] class_7.3-23 sass_0.4.10 yaml_2.3.12
#> [25] prodlim_2025.04.28 tidyr_1.3.2 pillar_1.11.1
#> [28] pkgdown_2.2.0 jquerylib_0.1.4 MASS_7.3-65
#> [31] cachem_1.1.0 gower_1.0.2 rpart_4.1.24
#> [34] nlme_3.1-168 parallelly_1.46.1 lava_1.8.2
#> [37] tidyselect_1.2.1 digest_0.6.39 future_1.69.0
#> [40] purrr_1.2.1 listenv_0.10.0 labeling_0.4.3
#> [43] splines_4.5.2 fastmap_1.2.0 grid_4.5.2
#> [46] cli_3.6.5 magrittr_2.0.4 utf8_1.2.6
#> [49] survival_3.8-3 future.apply_1.20.1 withr_3.0.2
#> [52] scales_1.4.0 lubridate_1.9.4 timechange_0.4.0
#> [55] rmarkdown_2.30 globals_0.19.0 nnet_7.3-20
#> [58] timeDate_4052.112 ragg_1.5.0 evaluate_1.0.5
#> [61] knitr_1.51 hardhat_1.4.2 mgcv_1.9-3
#> [64] rlang_1.1.7 Rcpp_1.1.1 glue_1.8.0
#> [67] ipred_0.9-15 jsonlite_2.0.0 R6_2.6.1
#> [70] systemfonts_1.3.1 fs_1.6.6