Overview
measure.sec is an R package that extends the measure package with preprocessing and analysis steps for Size Exclusion Chromatography (SEC) and Gel Permeation Chromatography (GPC) data.
Features
- Multi-detector support: RI, UV/DAD, MALS, LALS, RALS, DLS, and viscometer processing
- Inter-detector delay correction: Align signals from detectors in series
- Molecular weight calculations: Mn, Mw, Mz, dispersity (PDI)
- Copolymer composition: UV/RI ratio and composition analysis
- Protein SEC: Aggregate and fragment quantitation (HMWS/LMWS)
- Universal calibration: Mark-Houwink parameter-based MW conversion
- Quality control: System suitability testing (resolution, plate count, asymmetry)
- Polymer analysis: Branching indices and Mark-Houwink parameter estimation
Package Workflow
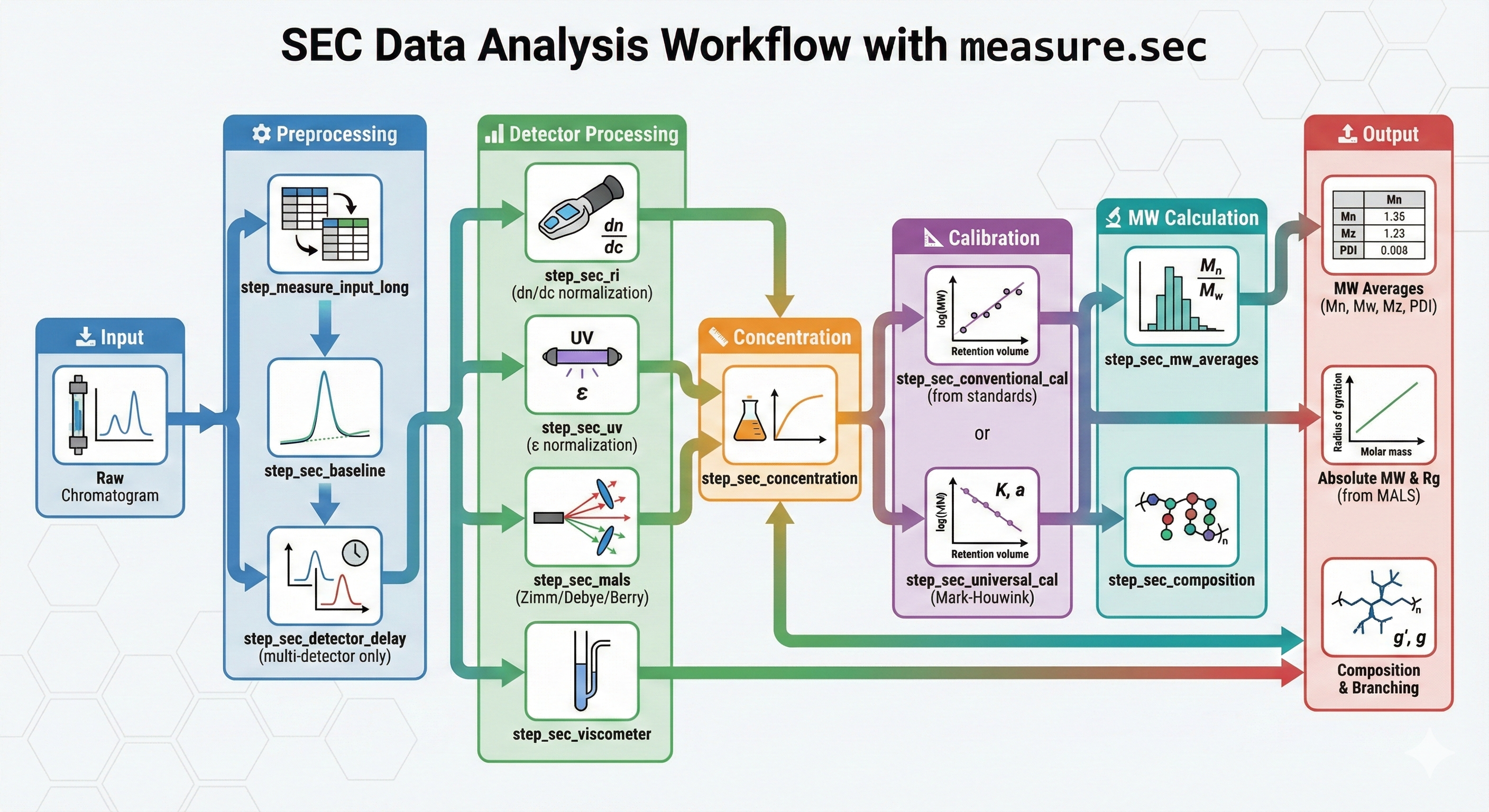
The typical SEC analysis workflow processes raw chromatogram data through preprocessing, detector-specific signal processing, calibration, and molecular weight calculations. Concentration detectors (RI, UV) require calibration curves from standards, while light scattering detectors (MALS) provide absolute molecular weight directly.
Installation
You can install the development version of measure.sec from GitHub with:
# install.packages("pak")
pak::pak("JamesHWade/measure-sec")Note: measure.sec requires the measure package:
pak::pak("JamesHWade/measure")Quick Start
library(measure)
library(measure.sec)
library(recipes)
# Load example triple-detection SEC data
data(sec_triple_detect, package = "measure.sec")
# Create a processing recipe
rec <- recipe(~ ., data = sec_triple_detect) |>
# Convert to measure format
step_measure_input_long(ri_signal, location = vars(elution_time), col_name = "ri") |>
step_measure_input_long(uv_signal, location = vars(elution_time), col_name = "uv") |>
# Correct inter-detector delays
step_sec_detector_delay(
reference = "ri",
delay_volumes = c(uv = -0.05)
) |>
# Apply baseline correction
step_sec_baseline() |>
# Process RI detector with dn/dc
step_sec_ri(dn_dc_column = "dn_dc")
# Prep and bake
prepped <- prep(rec)
result <- bake(prepped, new_data = NULL)Available Steps
Detector Processing
| Step | Description |
|---|---|
step_sec_ri() |
RI detector with dn/dc normalization |
step_sec_uv() |
UV detector with extinction coefficient |
step_sec_dad() |
Diode array detector processing (multi-wavelength UV) |
step_sec_mals() |
Multi-angle light scattering for absolute MW |
step_sec_lals() |
Low-angle light scattering for absolute MW |
step_sec_rals() |
Right-angle light scattering for absolute MW |
step_sec_dls() |
Dynamic light scattering for Rh and diffusion |
step_sec_viscometer() |
Differential viscometer processing |
step_sec_concentration() |
Convert signal to concentration |
step_sec_intrinsic_visc() |
Intrinsic viscosity calculation |
Preprocessing
| Step | Description |
|---|---|
step_sec_detector_delay() |
Correct inter-detector volume delays |
step_sec_baseline() |
SEC-optimized baseline correction |
Molecular Weight
| Step | Description |
|---|---|
step_sec_mw_averages() |
Calculate Mn, Mw, Mz, dispersity |
step_sec_mw_fractions() |
Calculate MW fractions above/below cutoffs |
step_sec_mw_distribution() |
Generate differential/cumulative MWD |
step_sec_conventional_cal() |
Narrow standard calibration (polynomial fits) |
step_sec_universal_cal() |
Universal calibration with Mark-Houwink |
Composition Analysis
| Step | Description |
|---|---|
step_sec_uv_ri_ratio() |
UV/RI ratio for heterogeneity detection |
step_sec_composition() |
Copolymer composition from UV/RI |
Protein SEC
| Step | Description |
|---|---|
step_sec_protein() |
Complete protein SEC workflow (baseline + aggregates + oligomers) |
step_sec_aggregates() |
HMWS/monomer/LMWS quantitation |
step_sec_oligomer() |
Detailed oligomer species analysis |
Quality Control Functions
# Peak resolution
Rs <- measure_sec_resolution(
retention_1 = 8.5,
retention_2 = 10.0,
width_1 = 0.3,
width_2 = 0.35
)
# Plate count
N <- measure_sec_plate_count(retention = 10.0, width = 0.25)
# System suitability testing
peaks <- data.frame(
name = c("dimer", "monomer"),
retention = c(8.5, 10.0),
width = c(0.3, 0.35),
area = c(5, 95)
)
sst <- measure_sec_suitability(
peaks = peaks,
reference_peaks = c("dimer", "monomer")
)
print(sst)Polymer Analysis Functions
# Estimate Mark-Houwink parameters
mw <- c(10000, 25000, 50000, 100000, 250000)
iv <- c(0.15, 0.28, 0.45, 0.72, 1.2)
mh <- measure_mh_parameters(mw, iv)
print(mh)
# K = 1.14e-04, a = 0.716
# Calculate branching index
g <- measure_branching_index(
mw = mw,
rg = c(8, 12, 18, 28, 45),
reference = data.frame(mw = mw, rg = c(10, 15, 22, 35, 55)),
method = "g"
)Data Export
# Extract slice-by-slice data
slices <- measure_sec_slice_table(result, measures = c("ri", "mw"))
# Generate summary table
summary <- measure_sec_summary_table(result, sample_id = "sample_id")Example Dataset
The package includes sec_triple_detect, a synthetic multi-detector SEC dataset with:
- 12 polymer samples (PS standards, PMMA, PEG, copolymers)
- RI, UV, and MALS detector signals
- Known molecular weights and dispersities
- Sample-specific dn/dc and extinction coefficients
Integration with measure
measure.sec automatically registers with the measure package on load:
library(measure)
library(measure.sec)
# View registered SEC steps
measure_steps(techniques = "SEC/GPC")Learn More
For detailed workflows and examples, see the package vignettes:
- Getting Started: Basic concepts and your first SEC analysis
- Multi-Detector SEC: Triple detection with RI, UV, and MALS
- Protein SEC: Aggregate and oligomer analysis for biopharmaceuticals
- Copolymer Analysis: Composition from UV/RI ratio
- System Suitability: QC functions and SST testing
Code of Conduct
Please note that the measure.sec project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.